2022.07.02.46
Files > Volume 7 > Vol 7 No 2 2022
1.2.3. Department of Microbiology, College of Medicine, Mustansiriyah University, Baghdad IRQA
*Corresponding author email is: [email protected],
Available from: http://dx.doi.org/10.21931/RB/2022.07.02.46
ABSTRACT
Escherichia coli are gram-negative bacteria that cause urinary tract infections (UTIs). UTIs have affected a significant percentage of humans yearly due to bacterial infection. Our study aims to determine the prevalence of resistance genes in E. coli towards sulfamethoxazole. This study included (490) patients with UTIs, and the urine samples were cultured on media. The patients were admitted to the Medical City in Baghdad to treat UTIs. 116 E.coli isolates were isolated from urine specimens, 35 isolates of them were resistant to trimethoprim/sulfamethoxazole, and 81 isolates were sensitive to trimethoprim/sulfamethoxazole; the E. coli isolates were submitted to multiplex PCR to detection some resistance genes (Sul1, sul2) after detected the isolates by PCR depending on 16S rRNA. Our study showed that identified E. coli was (91-99%) depending on the number of the examined samples by the Vitek 2 system. The molecular study included extraction of chromosomal DNA from (53) E. coli isolates; 35 samples were taken resistant to antibiotics, while from the total of 81 sensitive isolates, only 18 sensitive samples were taken from that are the most sensitive to Timethprime/sulfamethoxazole, then identification by 16S rRNA gene. Detection of Sulfonamides resistance genes included sul1 and sul2. The results showed the 16S rRNA gene identification found in all E. coli isolates and the detection of antibiotic resistance genes. The resistant isolates with the Sul1 gene prevalence were 11(31%), while the sensitive isolates with Sul1gene were 1(6%).
Moreover, the resisted isolates with Sul2 gene prevalence was 8(23%), while the sensitive isolates with the Sul1 gene were 0(0%). The numbers of the resistant isolates were (11) and (8) that carry the Sul1 gene and Sul2 gene, respectively, while the numbers of the sensitive isolates were (1) and (0), respectively. We can conclude that a high percentage of Sul1 gene and Sul2 genes in E. coil isolated from UTIs were high.
Keywords. UTI, Sul1, Sul2, resistant gene, trimethoprim-sulfamethoxazole
INTRODUCTION
Urinary Tract Infection (UTI) is a widespread disease in males and females. The occurrence percentage of Urinary Tract infections was 35% of healthy women with clinical signs of UTIs 1 . UTI was more incidence in females than males because of easy contamination with fecal flora, the squatness of female urethra dearth, and pregnancy 2. Many neonates, young females, infants, children and older men are infected with UTIs 3. Uncomplicated UTIs include bacterial entrance infection and bacterial proliferation in the urinary tract system 4. UTIs are mostly the second most common infections bacterial after respiratory tract infections. UTIs are inflammatory conditions in the urinary system leading to pyuria and bacteriuria 5. Antibiotic resistance has become a significant problem that needs coordinated action to reduce and prevent antibiotic resistance. The broad use of antibiotics for resistant bacteria may result in illness in humans that is less responsive to treatment with conventional antibiotics 6. Drug efflux pumps and the genes that respond to antibiotic resistance (by conjugation) are two mechanisms by which bacteria develop antibiotic resistance in large numbers 7. Trimethoprim and Sulphonamides are inexpensive antibiotics that work together to provide a synergistic effect. Since 1968, it has been used together as (co-trimoxazole) to treat some clinical cases, such as urinary tract infections. Trimethoprim sulphonamides resistance carries on the plasmids that generate the target enzymes that have a role in the resistance, such as dihydropteroate synthases against the sulphonamides and dihydrofolate reductases against the trimethoprim. Several genes, such as (sul1 and sul2) encode dihydropteroate synthases 8,9. Because of the importance and riskiness of E.coli, the increased incidence of infection and the possibility of the epidemic of infection as well as the riskiness of the disease and the lack of treatment, have led to the focus of research in the world on improving the efficiency of molecular detection using advanced technologies and reduce time and effort, the best diagnostic methods, such as PCR technique, are characterized by the technique of specificity and high speed in the detection of the genes encoding for the virulence and antibiotic resistance factors in isolates of E.coli isolated from the clinical specimens 10. This study aims to determine the prevalence and the resistance profile of E. coli in UTI, which resist Timethprime and sulfonamide such as sul1 and sul2 genes.
MATERIALS AND METHODS
Patient samples: This study included (490) patients suffering from urinary tract infections, the samples were acquired by culturing mid-stream urine.116 Escherichia coli sample isolate resistance trimethoprim-sulfamethoxazole 35 & sensitivity trimethoprim-sulfamethoxazole 81. 53 isolates were selected for genetic studies. The patients were admitted to the Medical City in Baghdad to treat UTIs between November 2020 and February 2021. Samples were collected in sterile cups using the mid-stream urine technique. The urine samples are collected into the sterile tubes; then, the urine is transported to the lab directly11.
Instruments and Equipment: Many instruments and many types of equipment have been used to process the collected samples, such as Autoclave, Centrifuge, Deep-freezer, Oven, Gel electrophoresis apparatus and power supply, Centrifuge, Eppendorf tube, PCR tube (0.2 µl), Incubator, Magpurix, Micro Centrifuge tubes, Micropipette, microscope, Nano-drop, Refrigerator, RT-PCR, Vortex mixer, Water bath.
The used Media: It is used throughout the study; the name of basic like nutrient agar & enrichment media.
The used primers: The source of all primers used in this study was Macrogen® (Korea). The name, sequence and product size are shown in table (1).

Table 1. The sequence of the used Primer with Product size
Isolation and Identification of Bacteria: Bacteria were isolated as pure colonies on MacConkey agar, Eosin methylene blue agar, blood agar, then bacterial isolates were examined and identified by microscopic, cultural, biochemical test and Vitek2 system 12.
Microscopical Examination (Gram stain): Bacterial isolates are examined for Gram stainability; shape and arrangement were observed 12. The appearance of colonies on the MacConkey agar and Eosin methylene blue agar and blood agar are studied concerning the shape, color and other characteristics 13. Some drops of 3% hydrogen peroxide reagent were added. The release of gas bubbles within (20–30) seconds indicated a positive result 14.
Confirmation of E. coli using API 20 E System: The API, 20 E strip system is a standardized identification technique for non-fastidious, enteric Gram-negative rods belonging to Enterobacteriaceae. The system comprises 20 microtubes that are packed along with dehydrated substrates. These tiny tubes contained the pure microbe that was incubated at (37) C for 24hour. The metabolism results in color changes; the test results are read, and the bacteria is detected. Each test's positive and negative findings are utilized to create a 7 code number to determine the microorganism identification.
Identification and Antibiotic Susceptibility Test
Genomic DNA extraction: gDNA was extracted from the blood according to the company's instructions.
Agarose Gel Electrophoresis: After extraction of gDNA, the electrophoresis is adopted to confirm the DNA 15.
The electrophoresis agarose gel: After sealing the tray's edges with adhesive tape and positioning the comb, the agarose is put into the gel tray at (18-22 C) for a half-hour, then removed from the comb; after that, the gel tray is put in the gel tank, which is filled with 1x TBE buffer until the gel tray is fully immersed 15.
DNA loading and electrophoresis: 7 DNA was combined with 3l of 6X loading dye. (Save stain) were put into the wells, then electrical power at 70V for half an hour, resulting in DNA migration from (-) to (+) poles. The save stain are stained bands were photographed and visualized using a UV light transilluminator device (Cleaver Scientific, USA)
Estimation of DNA concentration and purity: Nano-drop NAS-99 spectrophotometer was used to evaluate the concentration and purity of DNA in samples. The Nano-drop was first blanked with 2 µl of DNA rehydration solution (same elution material). The DNA samples were measured one by one, Estimating DNA concentration: reading A at 260nm.
Detection of 16s rRNA for E coli and sul1 gene And sul 2 gene: The primers in this study were used to amplify specific regions of the sul1 gene and sul2 gene. These primers were shown in (Table 2) as a lyophilized product of different Pico moles concentrations. According to the instructions of the manufacturing company, the primers are added to the water without nucleated at a concentration of 100 pmol /µl; from this solution (10 μl) was added to (90 μl) of water without nucleated to result in primer at (10 pmol/µl) as a table (2).
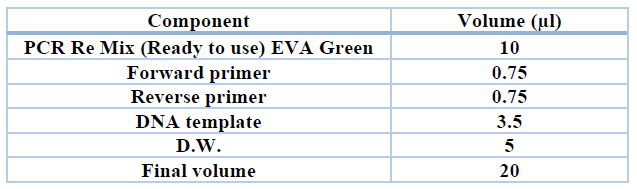
Table 2. Monoplex PCR reaction components for amplifying the targeted fragments to detect E coli ( 16 sRNA)
Polymerase chain Reaction components and programs: Polymerase – Chain reaction was carried out after several attempts of optimization to detect the best temperature for annealing with a total volume of 20 μl using LM 2012 Thermal cycler (Wizbio, South Korea). PCR Amplification Program is shown in table (3).

Table 3. Shows PCR Amplification Program
Dye detection (Filter HRM) for EVA green: HRM analysis is an effective method in molecular science for mutation detection, epigenetic and polymorphism differences in DNA samples 16.
RESULTS
Isolation of Escherichia coli: Four hundred ninety urine samples were collected from patients with a urinary tract infection (UTI). At the same time, 116 isolates were Escherichia coli, from many hospitals in Medical City in Baghdad (National Center for Educational Laboratories, Baghdad Teaching Hospital and Specialized Surgery Hospital) with different gender and ages, during the period between November 2020 and February 2021.
Identification of Escherichia coli
Microscopic examination: Escherichia coli isolates are examined under the light compound microscope shown as Gram-negative bacteria, rod-shaped, arranged in single or aggregated in pairs and non–spore-forming according to described 17.
Cultural Characteristics: Cultural characteristics for E. coli isolates appeared in the selective media. Morphology of the colony on the MacConkey are included large bright pink colonies due to lactose fermentation, circular, raised low convex with an entire edge, smooth surface and E. coli forming green metallic sheen colonies when grown on Eosin methylene blue.

Figure 1. Growth Culture for E. coli on (A) MacConkey agar and (B) blood agar showed colony morphology.
Biochemical Tests: Escherichia coli isolates give results the biochemical tests wherever, a positive result for catalase test, Indole test, Methyl red, and Motility, but they have given negative results to oxidase, Simmons Citrate, Voges Proskauer and Hydrogen Sulfide (H2S) production and give for Triple sugar agar test.
Identification using Vitek 2 system:
According to Table (4), the result shows that the identification probability of E. coli was (91-93%) for 8 isolates, while it was (93-96%) for 38 isolates and (96-99%) for 70 isolates by using the Vitek 2 system.

Table 4. Species identification Probability of isolates by the Vitek 2 system
Distribution of E. coli isolated according to hospital:
The overall number of collected specimens was 490, whereas the number of E. coli isolates was (116). The prevalence of the identified E. coli from National Center for Educational Laboratories was 75) 64.7 % (Baghdad Teaching was 29) 25 % (, and Specialized Surgery was 12)10.3% (, as shown in Table (5).

Table 5. Prevalence of E. coli isolates according to hospital
Escherichia coli antibiotic susceptibility:
The present study showed the percentage of resistance E. coli isolates against antibiotics as a table (6).

Table 6. Number and percentage of resistance, sensitivity, and intermediate of E. coli isolates
Genomic DNA Extraction: To genomic DNA was extracted from urine samples of patients and healthy controls by automated genomic DNA according to the standard protocol recommended by the manufactured company and manual extraction. Results of extraction showed that there are high concentrations and purity of DNA. The DNA concentration ranged between (110-120) ng/ µl, while the purity ranged between (1.8-2.0), calculated by the Nanodrop NAS-99 spectrophotometer. Genomic DNA extracted from each sample was analyzed on 1% agarose gel. Results of electrophoresis showed clear bands for genomic DNA extracted.
Relationship between antibiotics susceptibility and resistance genes:
Percentage of antibiotic resistant isolates and antibiotic sensitive isolates that carry the Sul1 gene and the Sul2 gene. Sul1 gene was found in 11(31%) of Trimethoprim/sulfamethoxazole resistant isolates. Sul2 gene was found in 8(23%) of Trimethoprim/sulfamethoxazole resistant isolates. The percentage of the antibiotics resistant isolates that have Sul1 gene was 11(31%), while the antibiotics sensitive isolates that have Sul1gene was 1(6%). The percentage of the antibiotics resistant isolates that have Sul2 gene was 8(23%), while the antibiotics sensitive isolates that have Sul1 gene was 0(0%), as shown in Table (7) and figure (2).

Table 7. Shows the number and percentage of resistance, sensitivity, and total isolates with Sul1 and SuL2 gene

Figure 2. Showed trimethoprim resistance genes wherever the horizontal line represents the temperature while the vertical line represents the df / dt
DISCUSSION
Distribution of age & sex among infected patients with E. coli Isolates
The current study found that the highest prevalence of E. coli isolates was at age (21-40) years in 15(51.7%) males and 40(46%) females, while the lowest prevalence was at age >=20 years at 1(3.4%) in the males and 12(13.8%) in the females. According to Al-Saadi (2018), distribution of the infected patients with four E. coli, three isolates was obtained from the female and one isolate from male according to within two age groups (41-50) and (61-70) years, respectively. Urinary tract infection was in age > 18 years was (12.9%) in females, while in the male (24.6%), the same study found that UTI was the percentage of the female at age (19–40) years was (32.4%), while the percentage of the male was (17. %) (16). the UTI percentage in males was (52%) while in females was (48%). Some reports were shown that the UTI rate was (23.3%) in males and was (76.7%) in females 18. While the previous stud showed that UTI rates were (44.24%) and (55.75%) in males and females, respectively 19. the percentage of UTs by E. coli was (35.3%) in males and (64.7%) in females20.
UTI Occurrence is associated with changes in the host's immunity, like immune suppression, and the diseases which cause immune inhibition such as hyperadrenocorticism, diabetes mellitus, tumors, uroliths, and indwelling catheters. E. coli are normal flora in the urethra and could be transmitted to all urinary tract parts. The virulence factors of E. coli help invade the cells and then generate toxins to inhibit immune system activity. Causes of UTI in females are common. Therefore it occurs in a high percentage. Short urethra and complex physiology during pregnancy are common causes of UTI in young females. Additionally, contraceptives could increase the risk factors of UTIs 7.
The highest percentage of isolates, 261(73.1%), were obtained from females and 96(26.7%) from males 21. The patient's age (2 months-90 years). The age (21-30) years group showed that E. coli was most common (23.53)%, followed by the group more than ten years (12.61)%, and the lowest rate was among the group (81-90) years (3.36)%. Eighty-one positive E. coli, the males were 17(21%) isolates, and the females were 64(79%) isolates. The age category (31-45) years is the most susceptible group for UTI was (41.98) %, followed by the age category (16-30) years was (27.16)%, the age category (46-60) years was (19.75)%, the age category (>60) years was 7.4% and the age category (0-15) years was (3.7)%. These results may suggest that UTI are common in the age category (16-60) years 22.
Antimicrobial Susceptibility:
The current study demonstrated that the percentage of resistance, sensitivity, and intermediate isolates toward the trimethoprim-sulfamethoxazole were 30.2%, 69.8%, and 0%, respectively. Many studies survey the percentage of Trimethoprim-sulfamethoxazole resistance in E. coli isolated from UTIs. The percentage of Trimethoprim-sulfamethoxazole's resistance was (91.1%) by 23, (26.9%) by 24, (32.25%) by 25, and (57.3%) by 26. In contrast, the prevalence of sensitivity to trimethoprim-sulfamethoxazole in E. coli isolates was (31.3%) by 27, (70.2%) by 28, (67.74%) by 24, and (20.0%) by 29. The resistance of bacteria to trimethoprim occurs due to higher production of the enzyme (DHFR) targeted promoter for the antibiotic by promoter mutation 29, and the resisted E. coli to Sulfonamides antibiotic by the massive production of the enzyme (PABA), the sulfonamides mimic PABA. Therefore, the antimicrobials have difficulty reaching the target 30. At the same time, the resistance of E. coli to Trimethoprim/sulfamethoxazole is due to chromosomal mutations in the dhfr or dhps genes that cause antibiotic resistance 31. Many studies have dealt with the issue of the emergence of the phenomenon of resistance to antibiotics by many bacteria. Antibiotic resistance is common among bacteria, especially bacteria that cause disease states and lead to treatment failure or unresponsiveness 32. Antibiotic resistance is activated by several mechanisms, either through genetic resistance, including the efflux mechanism or through acquired mechanisms and mutations in the genetic material and plasmids 33. There are six causes of antibiotic resistance: the high concentration of antibiotics, not finishing the therapeutics course, common use of antimicrobials in veterinary products, Poor control of the infection, and poor hygiene and sanitation. Antimicrobial resistance increases due to overuse and misuse of antibiotics and poor infection control. Therefore, the prevalence of antibiotic resistance depends on the risk factors. The absence of step factors or the presence of one or more of the above is the leading and only reason for the phenomenon of variation in the incidence of antibiotic resistance 34.
Prevalence of Sul1 gene in Tri / sulpha resistant E. coli isolates:
Sul1gene was (31%) in (trimethoprim/ sulfamethoxazole) resistant isolates. This is similar to other previous studies that found the distribution for the sul1gene in sulfamethoxazole-resistant isolates was 36.58%, 22.7%, 34%, 31.4%, and 32%, respectively 35-39. However, lower results were recorded, Sul1 was found in (10.3%) of sulfamethoxazole-resistant isolates (40), while higher prevalence was shown by previous studies, 53.0%, 45.9%, 41.4% 41-43. Cotrimoxazole Resistance sul1 Genes in E. coli isolates were 81.6 %. 44
Prevalence of Tri-sulpha resistant E. coli isolates with Sul2 gene:
The prevalence of the Sul2 gene was (23%) in studied E. coli. It is similar to what was found by 45, who reported that Sul2 genes and sulfamethoxazole resistance genes were found in the isolates at (18.5%). While it was lower than revealed by other studies, it reported that Sul2 genes, the most common sulfamethoxazole resistance gene, were (77.9%) isolates46. On the other hand, Cotrimoxazole Resistance sul 2 Genes in Escherichia coli Isolates (66.4 %) has appeared in another report43. Genes of sul 2 were found in 40 % of sulfamethoxazole-resistant isolates 45, but, it was shown (81.0%) in another article 46. The difference in the prevalence of resistance genes may be due to the difference between countries in the type of treatment used for urinary tract infections that might encourage antibiotic resistance mechanisms development due to high exposure for Trimethoprim/sulphamethoxazole, which is used as the first antibiotic against UTIs, the type of isolates present in each country and the rates of resistance genes in the bacterial isolates.
CONCLUSION
Escherichia coli isolates resistant to trimethoprim/sulfamethoxazole were high, and most of these isolates have a high percentage of resistance genes Sul1 gene and Sul2.
Acknowledgment: Thanks going for all who support us.
Conflict between authors: No conflict
Funds: self by authors
REFERENCES
1-Haque R, Akter ML, Salam MA. Prevalence and susceptibility of uropathogens: a recent report from a teaching hospital in Bangladesh. BMC Res Notes. 2015 Sep 5;8:416.
2- Nithyalakshmi J. Bacterial profile and antibiogram pattern of UTI in pregnant women at tertiary care teaching hospital. Int J Pharm Bio Sci 2014, 5: 201-207.
3- Yadav K, Prakash S, Serayi RC, Shilpkar T, Shrestha S. Antimicrobial susceptibility test of pathogens isolated from urinary tract infection suspected cases. Janaki Med Coll J Med Sci 2014, 2: 28-34.
4-Iroha I, Nwakeze E, Ejikeugwu C, Oji A, Udu-Ibiam E, Frequency and antibiogram of uropathogens isolated from urine samples of HIV infected patients on antiretroviral therapy. Am J Bio Sci 2013, 1: 50-53.
5-Forouzan MZA, Amir B. Prevalence and antimicrobial susceptibility patterns of uropathogens among patients referring to Valieasr laboratory in Najafabad, Isfahan, Iran. Middle-East J Sci Res 2013, 13: 85-90.
6-Roca, I., Akova, M., Baquero, F., Carlet, J., Cavaleri, M., Coenen, S., Heure, O. E. The global threat of antimicrobial resistance: science for intervention. New Microbes and New Infections, 2015, 6, 22–29.
7-Yadav K, Prakash S. Antimicrobial resistance (AMR): A global problem. Glob J Publ Health Epidemiol 2016, 3: 120-138.
8-Suzuki S, Hoa PT. Distribution of quinolones, sulfonamides, tetaracyclines in aquatic environment and antibiotic resistance in indochina. Front Microbiol. 2012;3:67.
9-Trobos M, Christensen H, Sunde M, Nordentoft S, Agerso Y, Si¬monsen GS. Characterization of sulphonamide-resistant Escherichia coli using comparison of sul2 gene sequences and multilocus sequence typing. Microbiology.; 2009, 155(Pt 3):831–6.
10-Fournier, P. E., Dubourg, G., & Raoult, D. Clinical detection and characterization of bacterial pathogens in the genomics era. Genome medicine, 2014, 6(11), 114.
11-Price, T. K., Dune, T., Hilt, E. E., Thomas-White, K. J., Kliethermes, S., Brincat, C., Brubaker, L., Wolfe, A. J., Mueller, E. R., & Schreckenberger, P. C. The Clinical Urine Culture: Enhanced Techniques Improve Detection of Clinically Relevant Microorganisms. Journal of clinical microbiology, 2016, 54(5), 1216–1222.
12-Benson, J. H. (2001). Microbiological Applications: Laboratory Manual in General Microbiology.8th ed. McGraw Hill, United States of America.pp:2001, 26
13-MacFaddin, J. F. Biochemical tests for identification of medical bacteria. 3rd ed, 2000.
14-Hussein, NH; Rasool, Kh.H. and Hussein, J.D. Frequency of Extended Spectrum Beta Lactamase producing Gram negative bacteria isolated from blood cultures at children hospital in Baghdad. I.J.S.R. 2015, 4(1): 10-13.
15-Sambrook, J. and Russel, D. W. Molecular cloning: A laboratory manual. (3rd ed). Cold Spring Harbor, USA. 2001, pp:5-52.
16-Wittwer CT. High‐resolution DNA melting analysis: advancements and limitations. Human mutation. 2009;30(6):857-9.
17-Ahmed Hossain ac Saeem, Arafat Hossaina Aneeka Nawar Fatemab Abrar, Wahabac Mohammad, Morshad AlamacMd. Nazrul Islame Mohammad, Zakir Hossaind Gias and U. Ahsana 2020.
18-Mostafa Boroumand ORCID, Mohsen Naghmachi and Mohammad Amin Ghatee, 2021. DOI : 10.5812/jjm.112547
19-Khushbu Yadav and Satyam Prakash. Department of Biochemistry, Janaki Medical College Teaching Hospital, Tribhuvan University, Janakpurdham, Nepal, 2017.
20-Ulhusa, Patil. and Ambala, Chaudhari. Optimal production of alkaline protease from solvent- tolerant alkalophilic Pseudomonas aeruginosa. MTCC 7926.India Journal of Biotechnology. 2011, 10: 329-339.
21- Siiri Koljalg ,Kai Truusalu, Inga Vainumäe, Jelena Stsepetova, Epp Sepp, and Marika Mikelssar. Journal of Clinical Microbiology 2020, Vol. 47, No. 1
22-Kerrn M T. Klemmensen, N. Frimodt-Møller, F. Espersen. Susceptibility of Danish Escherichia coli strains isolated from urinary tract infections and bacteraemia, and distribution of sul genes conferring sulphonamide resistance. Journal of antimicrobial chemotherapy. 2002 1;50(4):513-6. Elisabet Guiral. Jordi Boscha, Jordi Vila, b Sara M. Soto (2012).
23-Mohsin M. AL-Nasrawi , Ashwak B. AL-Hashimy. Iraqi Journal of Biotechnolo, 2020, Volume 19, Issue 3, Pages 42-48.
24-Virginia H. Fleming, BCPSa, Bryan White, Robin South wood, and BCPS CDE. The American Journal of Emergency Medicine 2014, Volume 32, Issue 8, August, Pages 864-870. Resistance of Escherichia coli urinary isolates in ED-treated patients from a community hospital
25-Baydaa H Abdullah, Dalia Abdalkader Shakur and Fitua Al-Saedi. Estimation of the Antibiotic'sbacterial sensitivity, resistance and intermediate resistance in patients with urinary, tract infection using VITEK 2 system, annals tropical medicine and public health, special issue. 2021, 24.
26- AL-Abidi, H. M. K. Isolation and Identification of The Aerobic Bacteria Causing Infection in AL-Diwaniya city. j.Al-qadisiyah. Pure. Scie. 2005, 14(2):1-10.
27-Sayed Nassereddin Mostafavi , Soodabeh Rostami ,Yasamin Rezaee Nejad , Behrooz Ataei ,and Sina Mobasherizadeh(2021). Arch Iran Med. 2021;24(3): 187-192.
28-Grakh K, Mittal D, Prakash A, Bangar YC. Assessing the potential risk factors associated with avian colibacillosis using a questionnaire survey, 2020.
29-Bodour Al-Assil, Maysa Mahfoud & Abdul Rezzak Hamzeh. First report on class 1 integrons and Trimethoprim-resistance genes from dfrA group in uropathogenic E. coli (UPEC) from the Aleppo area in Syria, Mob Genet Elements. 2013, 1;3(3):e25204.
30- Alekshun, M.N. and Levy, S.B. Molecular mechanisms of antibacterial multidrug resistance. Cell. 2007, 128:1037-1050.
31- Al-Hussaini, A.K.; Moharram, A.M.; Ismail, M.A. and Gharamah, A.A.Human microbial keratitis in Upper Egypt. J. Bas. App. Myco. Egypt.2010, 1: 1-10.
32- Al-Janabi, A. O. F.;Al-Ani, S. F. And Sarah, I. Biofilms Formation on Contact Lenses:Clinical and Bacteriological Study. Diy. J. M. 2013, 5(2):11-18.
33-Zhao , B.;Allinson , S. L.; Bentley, MA. A.;Martin ,A. J. and Fullwood , N. J.Targeted cornea limbal stem/progenitor cell transfection in an organ culture model. Investigative Ophthalmology & Visual Science, 2008.
34-Ventola C. L. The antibiotic resistance crisis: part 1: causes and threats. P & T : a peer-reviewed journal for formulary management, 2015, 40(4), 277–283.
35- Fazeli, N. and Momtaz, H. Virulence Gene Profiles of Multidrug-Resistant Pseudomonas aeruginosa Isolated from Iranian Hospital Infections. Iran. Red. Crescent. Med. J. 2014, 16(10):67-68.
36-Dadmanesh, M. ; Pilehvarzadeh, M.; Eramabadi, M.; Eramabadi, P.; Moghadam, M. B. And Mashayekhi, F. Community Acquired Pseudomonas aeroginosa Urinary Tract Infections in Children Hospitalized in a Baqiatallah Hospital, Tehran, Iran: Virulence Profile and Antibiotic Resistance Properties. Biosci. Biotech. Res. Asia. 2014, 11(2):417-426.
37- Ameer MA, Wasey A, Salen P. Escherichia Coli (E Coli 0157 H7). 2021, Available from: https://www.ncbi.nlm.nih.gov/books/NBK507845/
38- American Academy of Ophthalmology Cornea/External Disease Panel. Preferred Practice Pattern Guidelines.Bacterial keratitis–Limited Revision. San Francisco, CA: American Academy of Ophthalmology. (2011).
39- Aminov, R. I. Biotic acts of antibiotics. Front. Microbiol. 2013, 4:241-242.
40- Animesh, J.; Avinash, P.; Manav, Kh.; Subhadra, J.; Annie, M.; Rajeev, R. P.; Raja, N.; Savitri, Sh.; Taraprasad, D.; Harry, W. and Flynn, J.r.Combined ceftazidime and amikacin resistance among Gram-negative isolates in acute-onset postoperative endophthalmitis prevalence, antimicrobial susceptibilities, and visual acuity outcome, J. Ophth. Infla. Inf.3:5-6. Antimicrobial Agents and Chemotherapy 2013, Vol. 44, No. 4
41- Antunes, P., Machado, J., Sousa, J. C., & Peixe, L. Dissemination of sulfonamide resistance genes (sul1, sul2, and sul3) in Portuguese Salmonella enterica strains and relation with integrons. Antimicrobial agents and chemotherapy, 2005, 49(2), 836–839. https://doi.org/10.1128/AAC.49.2.836-839.2005
42-Ashaye, A.and Aimola, A.Keratitis in children as seen in a tertiary hospital in Africa. J Natl Med Assoc.2008, 100 (4):386-390.
43- Azghani, A. O. ;edinghaus,T. B R.; Klein, C.; et, al. Detection of elastase from Pseudomonas aeruginosa in sputum and its potential role in epithelial cell permeability. Lung.2000, 178:181-189.
44-Hadis Arabi, Iraj Pakzad, Ayat Nasrollahi, Hasan Hosainzadegan, Farid Azizi Jalilian, Morovvat Taherikalani, Naser Samadi, And Allireza Monadi Sefidan. Journal Listjundishapur J Microbiolv.8(7); 2015 Julpmc4584071
45-Blahna M. T., Zalewski C. A., Reuer J., Kahlmeter G., Foxman B., Marrs C. F. The role of horizontal gene transfer in the spread of trimethoprim-sulfamethoxazole resistance among uropathogenic Escherichia coli in Europe and Canada. J Antimicrob Chemother, 2006, 57:666–672 .
46- Baig, M. Sh.A.; Ali, M.A.; Khokar, A.R.and Ahmed, I. Pseudomonas endophthalmitis: An analysis of fifteen cases. Pakistan J. surg. 2008, 24(2):113-116.
Received: 17 January 2022 / Accepted: 29 March 2022 / Published:15 May 2022
Citation. Majeed Issa O, Abdul-Elah Bakir W, Ayad Abbas M. Laboratory diagnosis of urinary tract infections in patients with resistance genes towards antibiotics. Revis Bionatura 2022;7(2) 46. http://dx.doi.org/10.21931/RB/2022.07.02.46
