2021.06.02.31
Files > Volume 6 > Vol 6 No 4 2021 > Vol 6 No 2 2021
Major epigenetic factors associated with the novel coronavirus disease-2019 (COVID-19) severity

Ahmed A Mhawesh1, Daniah Muneam Hamid2, Abdolmajid Ghasemian3
Available from: http://dx.doi.org/10.21931/RB/2021.06.02.31
ABSTRACT
The worldwide spread and high rate of viral transmission and related morbidity and mortality of Coronavirus disease-19 (COVID-19) is a crisis. Some epigenetic determinants predispose individuals to severe infection. Patients with prior chronic medical illnesses (hypertension, diabetes, lupus, and chronic obstructive lung disease) are highly susceptible to the infection. The aging and diabetes pandemic possibly exacerbate the COVID-19 or SARS-CoV-2 pandemic by enhancing COVID-19 associated comorbidities. COVID-19 utilizes several proteins for tackling the host immune response associated with enhancing comorbidities. The angiotensin-converting enzyme (ACE) is a significant receptor for SARS-CoV-2, which significantly expresses higher among individuals with comorbidities and under stress conditions. Patients with systemic lupus erythematosus are also prone to be susceptible to the disease. Viral infections cause a defect in the DNA methylation in lupus, causing further ACE2 hypomethylation and overexpression, leading to viral binding and cytokine storm and tissue damage during COVID-19 infection. The microRNAs (miRNAs) epigenetics regulations also play a critical role in the suppression of immune responses.
Meanwhile, viral proteins interplays with the host cell are conferred primarily through TGF-β and HIF-1 signaling, endocytosis, autophagy, and Toll-like receptor signaling RIG-I signaling, Il-17 signaling, and fatty acid oxidation/degradation. Furthermore, the COVID19 patient's metabolic states determine the infection severity. Noticeably, ten human metabolic proteins, including SGTA, SPECC1, FGL2, PHB, STAT3, BCL2L1, CAV1, JUN, PPP1CA, and XPO1, interact with the SARSE-CoV-2. Interactions between SARSCoV's spike protein-containing lipid-rich membrane compartments and epigenetic modulations are considered targets to inhibit the viral infection. Therefore, it seems that epigenetics plays a substantial role in the COVID-19 severity. Future in-depth studies will be promising. Vaccine design, particularly regarding ACE viral receptor monoclonal antibodies, is a proposal alongside adhering to personal hygiene.
Keywords: Coronavirus disease-19, epigenetics, severe respiratory disease
INTRODUCTION
For fighting against every infection, a triangle including genetics, environment, and lifestyle is the primary determinant of triumph. The recent emergence and spread of novel Coronavirus disease known as COVID-19 or Severe Acute Respiratory Syndrome Coronavirus-2 (SARSE CoV-2, also coronavirus) in Wuhan in central China, has recently caused a pandemic scale of pneumonia in humans leading to concurrently and continuing high transmission, morbidity and mortality rat 1 Coronavirus subfamily is single-stranded positive-sense (+ssRNA) virus. The pandemic spread of novel coronavirus disease-19 (CoVID-19) in 2019-2020 originated from Wuhan, China, continues to affect human health and many life aspects and activities. The severity of infection increases with advancing age. Data suggests that very complex host-virus interplays occur during the SARS-CoV-2 infections (table1 and Figure 1).
Although the pathogenicity of SARSE-CoV-2 has not been entirely understood, extensive lung damage, enhanced infiltration of monocytes, macrophages, and neutrophils within the respiratory system and the blood storm of proinflammatory cytokines and chemokines are associated with the severity of infection 2, 3. Another mechanism of viral evasion includes delayed IFN type I transduction which stimulates the monocytes and macrophages and delays T-cells activation. Both the SARSE-CoV and SARSE-CoV-2 viruses bind to the angiotensin-converting enzyme receptor (ACE) via their spike protein receptor-binding domains which share 72% amino acid similarity 4, 5. Strikingly, the SARSE-CoV-2 domain has an incredibly higher receptor affinity. Higher expression of ACE among patients with comorbidities supposedly predisposes them to severe infection.
Moreover, it was stated that viral binding to the ACE downregulates its expression and leads to lower level biosynthesis of end-product vasodilator heptapeptide angiotensin 1-7. This, in turn, causes lung injury due to increased pulmonary vascular permeability. Another receptor for Coronaviruses includes a zinc peptidase known as aminopeptidase N (APN), which shares homology and membrane topological similarities with the ACE 4, 6, 7.
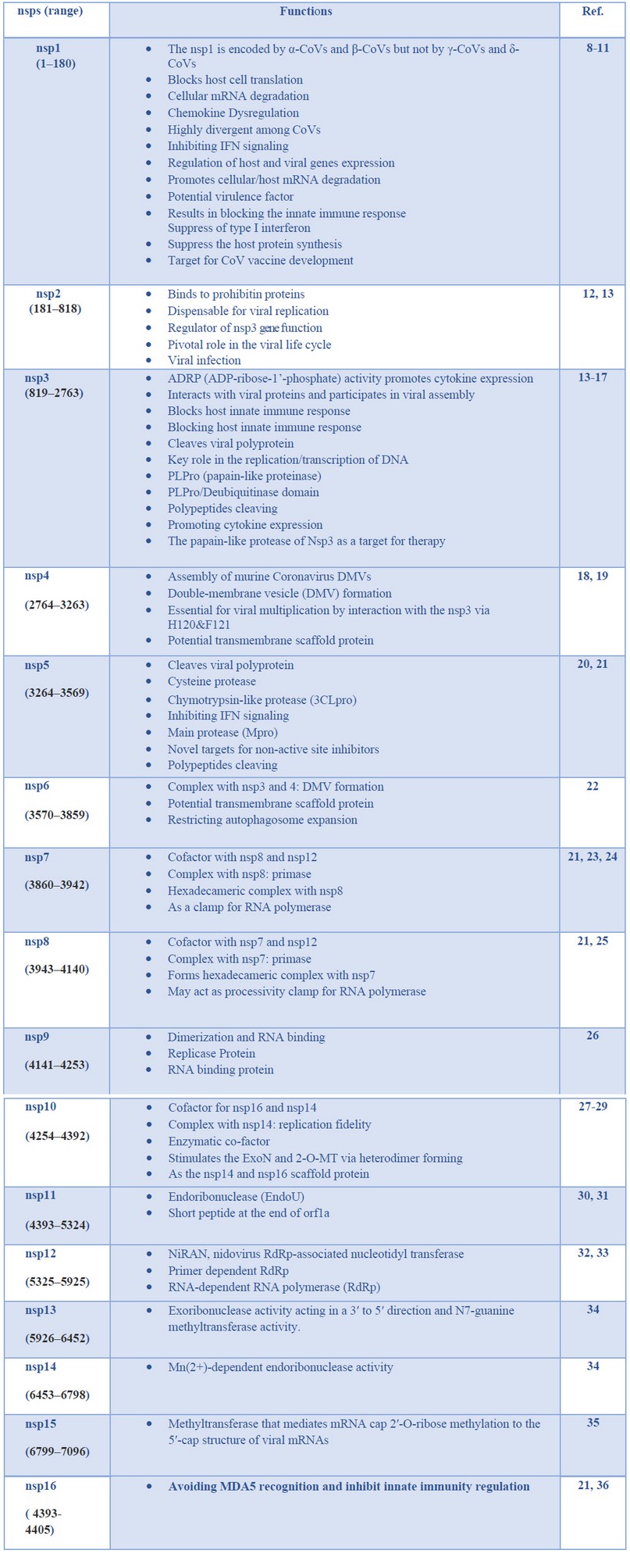
Table 1. Major Coronavirus non-structural proteins (nsps) and their pathogenicity capabilities
Those epigenetic factors mainly facilitating the viral attachment to host cells seem to enhance the death rate. These primarily include methylation or expression of angiotensin-converting enzyme 2 (ACE2), microRNAs regulation, metabolic conditions, individual behavior (such as smoking), and some environmental conditions (temperature and humidity) 37-39.
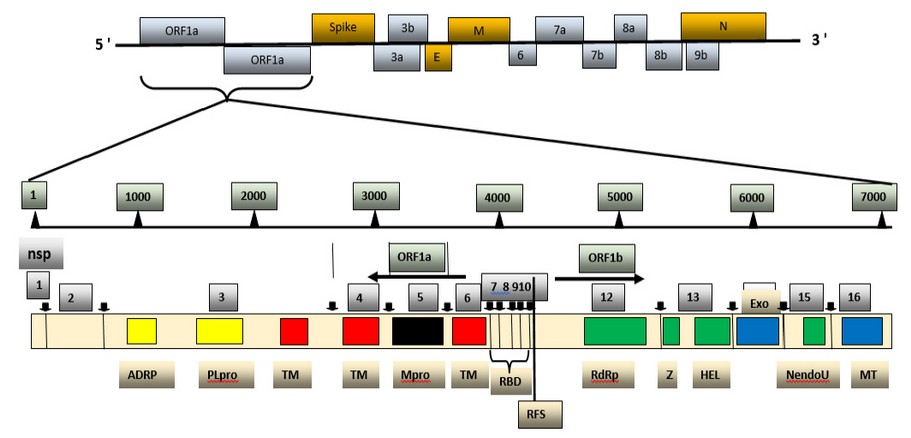
Figure 1. Genome and non-structural proteins of Severe Acute Respiratory Syndrome Coronavirus-2.
The genomic material released by its virus is mRNA, so it is prepared to stay translated into protein. In its genome range, its virus is complemented by using respecting 14 open reading frames (ORF), each of which encodes a variety concerning proteins, each structural yet non-structural, so move a function into its uplift so nicely as much virulence power. In its transformation section, the gene segments so encode non-structural polyproteins use this method to advance ORF1a yet ORF1b under production twins full-size overlapping polyproteins, pp1a and pp1ab utilizing contributing a ribosomal body shifting match 40 . The polyproteins are supplemented by using protease enzymes, specifically papain-like proteases (PLpro), yet a serine kind Mpro (chymotrypsin-like protease (3CLpro)) protease as are encoded of nsp3 then nsp 5. Subsequently, burst occurs into pp1a yet pp1ab into non-structural proteins (nsps) 1–11 and 1–16. The nsps shed a vital role in deep approaches of viruses yet host cells, as shown in Table 1. 2
Angiotensin-converting enzyme as the CoVID-19 receptor
Several body compartments, such as respiratory and gastrointestinal systems, have cells that express the ACE2 as the CoVID-19 receptor for its binding, entry (activation of the viral spike glycoprotein and ACE2 C-terminal segment cleavage), replication, and shedding. Epigenetics surveys have suggested that the ACE2 gene located on the X chromosome is regulated by DNA methylation 41-43. It was postulated that the variability in D/I genotype distribution of the ACE gene is possibly associated with the variable prevalence of the COVID-19 infection 42.
Notably, methylation varies across tissue cell types, and at three CpGs (cg04013915, cg08559914, and cg03536816) was lowest in lung epithelial cells. This figure was significantly lower among females than males, which differs in the ACE2 gene and protein expression and COVID-19 severity 44. Interestingly, increased ACE2 expression has been observed by staining lung tissue sections from patients with pulmonary hypertension. As the potential cause of ACE2 gene expression, enzymes that modify histones (KDM5B, H3K27a, H3K4me1, and H3K4me3) are notable. However, significant differences in ACE2 between racial groups, age groups, or gender groups were not verified in one study. Besides, smoking as an epigenome effector was not shown to have a role in COVID-19 infection risk. Also, IL-6 and INS (encoding the insulin hormone) genes have been associated with significant comorbidities. NAD-dependent histone deacetylase Sirtuin 1 (SIRT1) can also epigenetically induce the ACE gene expression under stress conditions. Viral infections cause a defect in the DNA methylation in lupus disease, causing further ACE2 gene hypomethylation and overexpression, leading to viral binding and cytokine storm and tissue damage COVID-19 infection 45, 46. The viral proteins exploit the host's genetic and epigenetic mediators, leading to viral evasion and determining disease pathophysiology (table1 and figure1). Besides, RAB1A gene was adequate for the COVID-19 infection development. Considering the ACE gene diffential expression, the age and sex of patients are also considered as risk factors for the COVID-19 infection 47.
Host and viral microRNAs epigenetic regulations
It has been revealed that epigenetics aspects of miRNA (small ncRNA molecules) mediated interactions with the host cells differ between SARSE-Cov and SARSE-Cov-2 (COVID-19). Hence, some viral miRNAs in a particular way affect several immune signaling pathways (IFN-I signaling, autophagy, etc.) that facilitate the prolonged latency. Moreover, COVID-19 modulates several critical cellular pathways resulting in the enhancement of anomalies in patients with comorbidities. The nucleocapsid protein from Coronavirus strain OC43 interacts with miR-9 and stimulates the NF-κB pathway. These findings are advantageous towards designing RNA therapeutics to mitigate the COVID-19 mediated comorbidities. Viral respiratory infections imposed by coronaviruses, influenza, adenovirus, rhinovirus, and RSV causes aberrant host miRNA expression mostly related to suppressing immune responses. Evidence has supposed that SARSE-Cov and SARSE-Cov-2 employ novel immune evasion strategies through utilizing host miRNA, but the exact mechanisms exerted by miRNA on the epigenetic interactions with the host have not been verified. It was hypothesized that genetic differences between SARSE-Cov and SARSE-Cov-2 and variations in binding to host miRNAs lead to differential pathogenesis 46.
Additionally, viral miRNAs differences and the fast mutation rate of SARSE-Cov2 in various regions have caused a higher pathogenicity rate. It was outlined that hsa-miR-20b-5p, hsa-miR-17-5p, and hsa-miR-323a-5p had anti-COVID-19 activities primarily targeting viral ORF1ab and S regions 44, 48. It is noteworthy that host miRNAs play a role like a double-edged sword and sometimes promote viral evasion, attachment, and replication. Patients suffering from underlying diseases (such as diabetes, cardiovascular diseases, and renal impairments) are more susceptible to SARS-CoV-2. It was revealed that host miRNA-mediated downregulated pathways disturb patients with the infection, making them more susceptible.
Viruses inflicting severe pulmonary illness can use three epigenetic-regulated approaches in the course of host-pathogen interaction: i. they execute affect host DNA methylation signatures then miRNAs regulating a cassette of genes underlying native yet adaptive antiviral responses; ii. those can encode because viral proteins up to expectation at once interact along with the host modified histones. Yet, iii. that may manipulate the host miRNA technology nuclear equipment in imitation of encoding viral non-canonical miRNA-like RNA fragments (v-miRNAs) regulating the viral life cycle or immune response 49. Here, we center on epigenetic-sensitive mechanisms via who H5N1 and SARS-CoV-2 can also affect susceptibility in imitation of pulmonary sickness by interfering with both born yet adaptive immune responses into human beings, as shown in Figure 2. 50, 51.
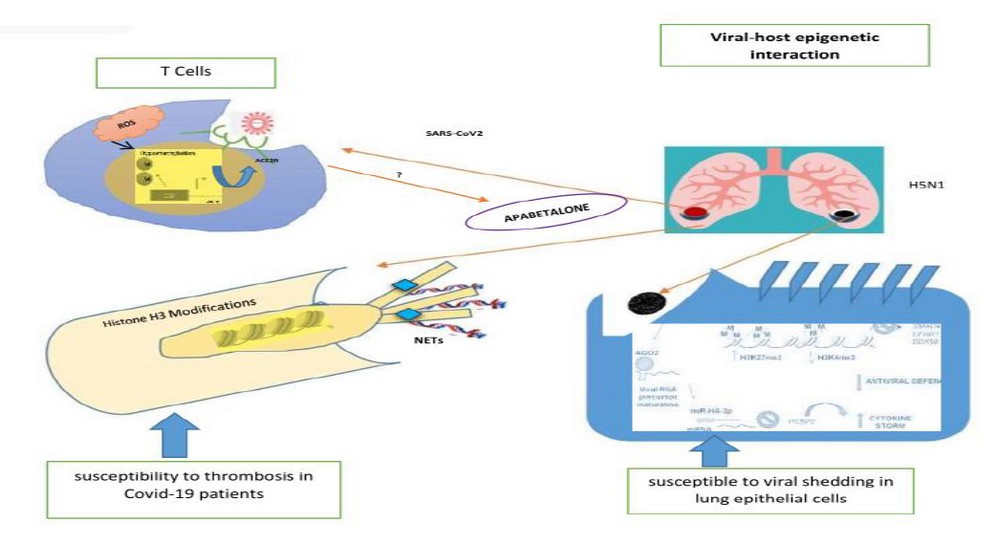
Figure 2. Viral–host epigenetic interactions. We illustrated that III putative cell-specific epigenetic-sensitive mechanisms using SARS-CoV-2 or H5N1 might also affect single sensitiveness according to severe pulmonary quintessential illness.
Mainly, T cells yet neutrophils execute bear deoxyribonucleic acid hypomethylation yet histone modifications, respectively, in COVID-19 patients. Otherwise, in vitro lung epithelial cells above H5N1 contamination can undergo adjustments of micro-ribonucleic sour taste (RNA) patterns yet histone tail marks, propulsion according to downregulation regarding antiviral defense. ACE, angiotensin-converting enzyme; AGO2, argonaute 2; CGI, CpG island; COVID-19, coronavirus disorder 2019; mRNA, messenger RNA; NET, neutrophil extracellular trap; NS1, non-structural protein 1; PCBP2, poly (RC)-binding protein 2; ROS, reactive oxygen species; SARS-CoV-2, severe acute respiratory indication coronavirus 2; SLE, systemic lupus erythematosus.
Patients'behavior, nutrition, and metabolic conditions
COVID19 patient's metabolic states determine the infection severity. Noticeably, ten human proteins, including SGTA, SPECC1, PHB, BCL2L1, FGL2, STAT3, JUN, PPP1CA, CAV1, and XPO1 interact with the SARSE-CoV-2. Interactions between SARSCoV's spike protein with lipid-rich membrane compartments and epigenetic modulations are considered targets to inhibit the viral infection 52. It was revealed that obesity is a risk factor for the SARSE-CoV-2 infection 53. Noticeably, ten human metabolic proteins, including SGTA, SPECC1, PHB, BCL2L1, FGL2, STAT3, JUN, PPP1CA, CAV1, and XPO1 interact with the SARSE-CoV-2. Interactions between SARSCoV's spike proteins, which have membranes rich in lipids components and epigenetic modulations, are considered targets to inhibit the viral infection. Some personal behaviors such as smoking or common personal behaviors that can reduce COVID-19 exposure are also notable, affecting the transmission, morbidity, and mortality rate 54, 55.
Future perspectives
Owing to the lack of effective vaccination until now, investigations in this regard will be promising. Additionally, the application of lower toxic disinfectants and adherence to personal hygiene; particularly among individuals meeting healthcare and developing effective drugs, can be helpful towards reducing the burden of infection. More in-depth verifications regarding epigenetic factors and application inhibitors of the ACE and outcomes are also proposed
CONCLUSION
Those epigenetic factors mainly facilitating the viral attachment to host cells seem to enhance the death rate. These primarily include methylation or expression of angiotensin-converting enzyme 2 (ACE2), microRNAs regulation, metabolic conditions, individual behavior (such as smoking), and some environmental conditions (temperature and humidity). The angiotensin-converting enzyme (ACE) is a significant receptor for SARS-CoV-2, which significantly expresses higher among individuals with comorbidities and under stress conditions. Patients with systemic lupus erythematosus are also prone to be susceptible to the disease. Viral infections cause a defect in the DNA methylation in lupus, causing further ACE2 hypomethylation and overexpression, leading to viral binding and also cytokine storm and tissue damage during COVID-19 infection. The microRNAs (miRNAs) epigenetics regulations also play a critical role in the suppression of immune responses.
Meanwhile, viral proteins interplays with the host cell are conferred mainly through TGF-β and HIF-1 signaling, endocytosis, autophagy, and Toll-like receptor signaling RIG-I signaling, Il-17 signaling, and fatty acid oxidation/degradation. Furthermore, the COVID19 patient's metabolic states determine the infection severity. Noticeably, ten human metabolic proteins, including SGTA, SPECC1, PHB, BCL2L1, FGL2, STAT3, JUN, PPP1CA, CAV1, and XPO1 interact with the SARSE-CoV-2. Interactions between SARSCoV's spike structural proteins containing membranes rich in lipidic macromolecules and epigenetic modulations are considered targets to inhibit the viral infection. Therefore, it seems that epigenetics plays a substantial role in the COVID-19 severity. Future in-depth studies will be promising. Vaccine design, particularly regarding ACE viral receptor monoclonal antibodies, is a proposal alongside adhering to personal hygiene.
Conflict of interest
None to declare
Acknowledgments
The authors wrote this study.
REFERENCES
1. Novel, CPERE, 2020. The epidemiological characteristics of an outbreak of 2019 novel coronavirus diseases (COVID-19) in China. Zhonghua liu xing bing xue za zhi= Zhonghua liuxingbingxue zazhi, 41(2), p.145.
2. Xu, Z., Shi, L., Wang, Y., Zhang, J., Huang, L., Zhang, C., Liu, S., Zhao, P., Liu, H., Zhu, L. and Tai, Y., 2020. Pathological findings of COVID-19 associated with acute respiratory distress syndrome. The Lancet respiratory medicine, 8(4), pp.420-422.
3. Rothan, H.A. and Byrareddy, S.N., 2020. The epidemiology and pathogenesis of coronavirus disease (COVID-19) outbreak. Journal of autoimmunity, p.102433.
4. Patel, A.B. and Verma, A., 2020. COVID-19 and angiotensin-converting enzyme inhibitors and angiotensin receptor blockers: what is the evidence? Jama, 323(18), pp.1769-1770.
5. Vaduganathan, M., Vardeny, O., Michel, T., McMurray, J.J., Pfeffer, M.A. and Solomon, S.D., 2020. Renin–angiotensin–aldosterone system inhibitors in patients with Covid-19. New England Journal of Medicine, 382(17), pp.1653-1659.
6. Fang, L., Karakiulakis, G. and Roth, M., 2020. Are patients with hypertension and diabetes mellitus at increased risk for COVID-19 infection? The Lancet. Respiratory Medicine, 8(4), p. e21.
7. Diaz, J.H., 2020. Hypothesis: angiotensin-converting enzyme inhibitors and angiotensin receptor blockers may increase the risk of severe COVID-19. Journal of Travel Medicine.
8. Narayanan, K., Ramirez, S.I., Lokugamage, K.G. and Makino, S., 2015. Coronavirus non-structural protein 1: Common and distinct functions in the regulation of host and viral gene expression. Virus research, 202, pp.89-100.
9. HUI, WH, 2012. Host-viral Interactions: Host factors in coronavirus replication and coronaviral strategies of immune evasion (Doctoral dissertation).
10. Sun, L., Xing, Y., Chen, X., Zheng, Y., Yang, Y., Nichols, D.B., Clementz, M.A., Banach, B.S., Li, K., Baker, S.C. and Chen, Z., 2012. Coronavirus papain-like proteases negatively regulate antiviral innate immune response through disruption of STING-mediated signaling. PloS one, 7(2), p. e30802.
11. Kamitani, W., Narayanan, K., Huang, C., Lokugamage, K., Ikegami, T., Ito, N., Kubo, H. and Makino, S., 2006. Severe acute respiratory syndrome coronavirus nsp1 protein suppresses host gene expression by promoting host mRNA degradation. Proceedings of the National Academy of Sciences, 103(34), pp.12885-12890.
12. Graham, R.L., Sims, AC, Baric, RS and Denison, M.R., 2006. The nsp2 proteins of mouse hepatitis virus and SARS coronavirus are dispensable for viral replication. In The Nidoviruses (pp. 67-72). Springer, Boston, MA.
13. Fehr, A.R. and Perlman, S., 2015. Coronaviruses: an overview of their replication and pathogenesis. In Coronaviruses (pp. 1-23). Humana Press, New York, NY.
14. Fehr, A.R., Athmer, J., Channappanavar, R., Phillips, J.M., Meyerholz, D.K. and Perlman, S., 2015. The nsp3 macrodomain promotes virulence in mice with coronavirus-induced encephalitis. Journal of virology, 89(3), pp.1523-1536.
15. Imbert, I., Snijder, E.J., Dimitrova, M., Guillemot, J.C., Lécine, P. and Canard, B., 2008. The SARS-Coronavirus PLnc domain of nsp3 as a replication/transcription scaffolding protein. Virus research, 133(2), pp.136-148.
16. Clementz, M.A., Kanjanahaluethai, A., O'Brien, T.E. and Baker, S.C., 2008. Mutation in murine coronavirus replication protein nsp4 alters assembly of double membrane vesicles. Virology, 375(1), pp.118-129.
17. Lei, J., Kusov, Y. and Hilgenfeld, R., 2018. Nsp3 of coronaviruses: Structures and functions of a large multi-domain protein. Antiviral research, 149, pp.58-74.
18. Sakai, Y., Kawachi, K., Terada, Y., Omori, H., Matsuura, Y. and Kamitani, W., 2017. Two-amino acids change in the nsp4 of SARS coronavirus abolishes viral replication. Virology, 510, pp.165-174.
19. Tomar, S., Johnston, M.L., John, S.E.S., Osswald, H.L., Nyalapatla, P.R., Paul, L.N., Ghosh, A.K., Denison, M.R. and Mesecar, AD, 2015. Ligand-induced dimerization of middle east respiratory syndrome (MERS) coronavirus nsp5 protease (3CLpro) implications for nsp5 Regulation and The Development of Antivirals. Journal of Biological Chemistry, 290(32), pp.19403-19422.
20. Qiu, Y. and Xu, K., 2020. Functional studies of the coronavirus non-structural proteins. STEMedicine, 1(2), pp. e39-e39.
21. Chen, Y., Liu, Q. and Guo, D., 2020. Emerging coronaviruses: genome structure, replication, and pathogenesis. Journal of medical virology, 92(4), pp.418-423.
22. Alexander, S.P., Ball, J.K. and Tsoleridis, T., 2020. SARS-CoV-2 proteins (version 2020.2) in the IUPHAR/BPS Guide to Pharmacology Database. IUPHAR/BPS Guide to Pharmacology CITE, 2020(2).
23. Subissi, L., Posthuma, C.C., Collet, A., Zevenhoven-Dobbe, J.C., Gorbalenya, A.E., Decroly, E., Snijder, E.J., Canard, B. and Imbert, I., 2014. One severe acute respiratory syndrome coronavirus protein complex integrates processive RNA polymerase and exonuclease activities. Proceedings of the National Academy of Sciences, 111(37), pp. E3900-E3909.
24. Zhai, Y., Sun, F., Li, X., Pang, H., Xu, X., Bartlam, M. and Rao, Z., 2005. Insights into SARS-CoV transcription and replication from the structure of the nsp7–nsp8 hexadecamer. Nature structural & molecular biology, 12(11), pp.980-986.
25. Egloff, M.P., Ferron, F., Campanacci, V., Longhi, S., Rancurel, C., Dutartre, H., Snijder, E.J., Gorbalenya, A.E., Cambillau, C. and Canard, B., 2004. The severe acute respiratory syndrome-coronavirus replicative protein nsp9 is a single-stranded RNA-binding subunit unique in the RNA virus world. Proceedings of the National Academy of Sciences, 101(11), pp.3792-3796.
26. Klosterman, S.J., Subbarao, K.V., Kang, S., Veronese, P., Gold, S.E., Thomma, B.P., Chen, Z., Henrissat, B., Lee, Y.H., Park, J. and Garcia-Pedrajas, M.D., 2011. Comparative genomics yields insights into niche adaptation of plant vascular wilt pathogens. PLoS pathog, 7(7), p. e1002137.
27. Bouvet, M., Lugari, A., Posthuma, C.C., Zevenhoven, J.C., Bernard, S., Betzi, S., Imbert, I., Canard, B., Guillemot, J.C., Lécine, P. and Pfefferle, S., 2014. Coronavirus Nsp10, a critical co-factor for activation of multiple replicative enzymes. Journal of Biological Chemistry, 289(37), pp.25783-25796.
28. Ma, Y., Wu, L., Shaw, N., Gao, Y., Wang, J., Sun, Y., Lou, Z., Yan, L., Zhang, R. and Rao, Z., 2015. Structural basis and functional analysis of the SARS coronavirus nsp14–nsp10 complex. Proceedings of the National Academy of Sciences, 112(30), pp.9436-9441.
29. Deng, X. and Baker, S.C., 2018. An "Old" protein with a new story: Coronavirus endoribonuclease is important for evading host antiviral defenses. Virology, 517, pp.157-163.
30. Woo, P.C., Huang, Y., Lau, S.K., Tsoi, H.W. and Yuen, K.Y., 2005. In silico analysis of ORF1ab in coronavirus HKU1 genome reveals a unique putative cleavage site of coronavirus HKU1 3C‐like protease. Microbiology and immunology, 49(10), pp.899-908.
31. Posthuma, C.C., te Velthuis, A.J. and Snijder, E.J., 2017. Nidovirus RNA polymerases: complex enzymes handling exceptional RNA genomes. Virus research, 234, pp.58-73.
32. Snijder, E.J., Bredenbeek, P.J., Dobbe, J.C., Thiel, V., Ziebuhr, J., Poon, L.L., Guan, Y., Rozanov, M., Spaan, W.J. and Gorbalenya, A.E., 2003. Unique and conserved features of genome and proteome of SARS-coronavirus, an early split-off from the coronavirus group 2 lineage. Journal of molecular biology, 331(5), pp.991-1004.
33. Ahn, D.G., Choi, J.K., Taylor, D.R. and Oh, J.W., 2012. Biochemical characterization of a recombinant SARS coronavirus nsp12 RNA-dependent RNA polymerase capable of copying viral RNA templates. Archives of virology, 157(11), pp.2095-2104.
34. Ahmad Abu TurabNaqviaKisaFatimabTajMohammadaUroojFatimacIndrakant K. SinghdArchanaSingheShaikh MuhammadAtiffGururaoHariprasadgGulam MustafaHasanhMd. ImtaiyazHassana, 2020. Insights into SARS-CoV-2 genome, structure, evolution, pathogenesis and therapies: Structural genomics approach. Biochimica et Biophysica Acta (BBA) - Molecular Basis of Disease.Vol. 1866, Issue 10, 165878.
35. M. von Grotthuss, L.S. Wyrwicz, L. Rychlewski.mRNA cap-1 methyltransferase in the SARS genome, 2003.Cell, 113, pp. 701-702.
36. Shi P, Su Y, Li R, Liang Z, Dong S, Huang J., 2019. PEDV nsp16 negatively regulates innate immunity to promote viral proliferation. Virus Res.; 265:57‐66.
37. Brake, SJ, Barnsley, K., Lu, W., McAlinden, K.D., Eapen, MS and Sohal, SS, 2020. Smoking upregulates angiotensin-converting enzyme-2 receptor: a potential adhesion site for novel coronavirus SARS-CoV-2 (Covid-19).
38. Zhang, P., Zhu, L., Cai, J., Lei, F., Qin, J.J., Xie, J., Liu, Y.M., Zhao, Y.C., Huang, X., Lin, L. and Xia, M., 2020. Association of inpatient use of angiotensin converting enzyme inhibitors and angiotensin II receptor blockers with mortality among patients with hypertension hospitalized with COVID-19. Circulation research.
39. Wang, J., Tang, K., Feng, K. and Lv, W., 2020. High temperature and high humidity reduce the transmission of COVID-19. Available at SSRN 3551767.
40. Masters P.S. The molecular biology of coronaviruses. Adv Virus Res., 2006; 66:193–292.
41. Crackower, M.A., Sarao, R., Oudit, G.Y., Yagil, C., Kozieradzki, I., Scanga, S.E., Oliveira-Dos-Santos, A.J., da Costa, J., Zhang, L., Pei, Y. and Scholey, J., 2002. Angiotensin-converting enzyme 2 is an essential regulator of heart function. Nature, 417(6891), pp.822-828.
42. Delanghe, J., Speeckaert, M. and De Buyzere, M., 2020. The host's angiotensin-converting enzyme polymorphism may explain epidemiological findings in COVID-19 infections. Clinica Chemica Acta.
43. Rasmussen, E.R., Hallberg, P., Baranova, E.V., Eriksson, N., Karawajczyk, M., Johansson, C., Cavalli, M., Maroteau, C., Veluchamy, A., Islander, G. and Hugosson, S., 2020. Genome-wide association study of angioedema induced by angiotensin-converting enzyme inhibitor and angiotensin receptor blocker treatment. The Pharmacogenomics Journal, pp.1-14.
44. Corley, M.J. and Ndhlovu, L.C., 2020. DNA methylation analysis of the COVID-19 host cell receptor, angiotensin I converting enzyme 2 gene (ACE2) in the respiratory system reveal age and gender differences.
45. Pinto, B.G., Oliveira, A.E., Singh, Y., Jimenez, L., Gonçalves, A.N.A., Ogava, R.L., Creighton, R., Peron, J.P.S. and Nakaya, H.I., 2020. ACE2 expression is increased in the lungs of patients with comorbidities associated with severe COVID-19. MedRxiv.
46. Sawalha, A.H., Zhao, M., Coit, P. and Lu, Q., 2020. Epigenetic dysregulation of ACE2 and interferon-regulated genes might suggest increased COVID-19 susceptibility and severity in lupus patients. Clinical Immunology, p.108410.
47. Chai, P., Yu, J., Ge, S., Jia, R. and Fan, X., 2020. Genetic alteration, RNA expression, and DNA methylation profiling of coronavirus disease 2019 (COVID-19) receptor ACE2 in malignancies: a pan-cancer analysis. Journal of Hematology & Oncology, 13, pp.1-5.
48. Khan, M.A.A.K., Sany, M.R.U., Islam, M.S., Mehebub, M.S. and Islam, A.B., 2020. Epigenetic regulator miRNA pattern differences among SARS-CoV, SARS-CoV-2 and SARS-CoV-2 worldwide isolates delineated the mystery behind the epic pathogenicity and distinct clinical characteristics of pandemic COVID-19. bioRxiv.
49. Gómez-Díaz E., Jordà M., Peinado M.A., Rivero A. Epigenetics of host–pathogen interactions: the road ahead and the road behind. PLoS Pathog. 2012;8
50. Iwasaki A., Foxman E., Molony R. Early local immune defences in the respiratory tract. Nat Rev Immunol.2017; 17:7–20.
51. Chiu C., Openshaw P. Antiviral B cell and T cell immunity in the lungs. Nat Immunol.2015; 16:18–26.
52. Kamepalli, M.D. and FIDSA, C., 2020. How Immune T-Cell Augmentation Can Help Prevent COVID-19: A Possible Nutritional Solution Using Ketogenic Lifestyle. The University of Louisville Journal of Respiratory Infections, 4(1), p.7.
53. Petrakis, D., Margină, D., Tsarouhas, K., Tekos, F., Stan, M., Nikitovic, D., Kouretas, D., Spandidos, D.A. and Tsatsakis, A., 2020. Obesity‑a risk factor for increased COVID‑19 prevalence, severity and lethality. Molecular Medicine Reports, 22(1), pp.9-19.
54. Basch, C.H., Hillyer, G.C., Meleo-Erwin, Z.C., Jaime, C., Mohlman, J. and Basch, C.E., 2020. Preventive behaviors conveyed on YouTube to mitigate transmission of COVID-19: cross-sectional study. JMIR public health and surveillance, 6(2), p. e18807.
55. Vardavas, C.I. and Nikitara, K., 2020. COVID-19 and smoking: A systematic review of the evidence. Tobacco induced diseases, 18.
Received: 24 January 2021
Accepted: 12 March 2021
Ahmed A Mhawesh1, Daniah Muneam Hamid2, Abdolmajid Ghasemian3
1Dept. of Med. and Mol. Biotech., College of Biotechnology, Alnahrain Univesirt, Baghdad, Iraq
2DNA Forensic center for research and training, Alnahrain University, Baghdad, Iraq
3 Islamic Azad University, Central Tehran Branch/Tehran/Iran
Corresponding author: [email protected]
1 Orcid ID: https://orcid.org/0000-0003-4546-7264
2 https://orcid.org/0000-0001-5539-3399
3 https://orcid.org/0000-0002-1243-6341