2021.06.04.7
Files > Volume 6 > Vol 6 No 4 2021
INVESTIGATION / RESEARCH
Expression of the ANS, CHS and DFR genes involved in the biosynthesis of anthocyanins in Vaccinium floribundum Kunth from Ecuador, using RT-qPCR.
Viviana Chiluisa-Utreras1*, Doris Vela2, Ivonne Vaca1, Ramiro Acurio1, Javier Chicaiza3 y Sofía Peñaherrera3.
Available from: http://dx.doi.org/10.21931/RB/2021.06.04.7
ABSTRACT
Vaccinium floribundum Kunth, a wild native species of berry in Ecuador, presents a lot of phenolic compounds, specifically anthocyanins; hence it is considered a natural nutraceutical due to all its nutritional properties. The comparison of the expression of genes is involved in the biosynthesis pathway of anthocyanin of several populations. The aim of the research was to analyze the expression levels of three genes involved in the biosynthesis of anthocyanin in this species collected in two areas of the province of Pichincha: Machachi population of the Mejía canton, with geographic coordinates 0 ° 31′04.8 ″ S 78 ° 37′07.4 ″ W and altitude 3200 meters above sea level, and Pintag population of the Quito cantón, with geographic coordinates 0 ° 24′00.0 ″ S 78 ° 24′00.0 ″ W and altitude 3000 meters above sea level. The gene expression analysis was performed using the quantitative polymerase chain reaction technique and reverse transcription (RT-qPCR). For the population of Machachi, the glyceraldehyde-3-phosphate dehydrogenase gene had an average concentration of 648.59 ng/µL, followed by the chalcone synthase gene with 143.71 ng/µL, then by the dihydroflavonol 4-reductase gene with 59.58 ng/µL and finally by the anthocyanin synthase gene with 39 ng/µL. For the population of Pintag, the glyceraldehyde-3-phosphate dehydrogenase gene has an average concentration of 667.32 ng/µL, followed by the chalcone synthase gene with 157.22 ng/µL, then by the dihydroflavonol 4-reductase gene with 60.42 ng/µL, and finally by the anthocyanin synthase gene with 44.40 ng/µL. Each gene has a similar expression level in both populations, but there are differences when comparing the expression level among genes. Many enzymes, structural genes, and regulatory elements have been observed as transcription factors involved in anthocyanin biosynthesis.
Key words: Anthocyanin synthase, chalcone synthase, dihydroflavonol 4-reductase, glyceraldehyde-3-phosphate dehydrogenase, Mortiño.
INTRODUCTION
The Vaccinium floribundum Kunth, known as Mortiño in Ecuador, is an endemic species from the north of South America between Colombia and Ecuador and native of the Ecuadorian Moors1. There are no commercial crops of this species, but only parcels of land where the species grows wildly, and its fruits are sold in the last months of the year2. In Ecuador, it is usually consumed as fresh fruit or it is processed in artisanal jams. The consumption of the fresh fruit is intended for preparing the traditional "colada Morada," a typical drink made in Ecuador at the October and November.
Besides being a fruit of great importance for the conservation of the moorlands and culture of Ecuador, Mortiño is a wild shrub still in the process of domestication in the country but with slow development and requires more in-depth studies. It possesses excellent nutritional properties related to its high content of phenolic compounds: anthocyanin3.
Anthocyanin is a type of flavonoid mainly involved in fruit ripening. These phenolic compounds are found in leaves, flowers, and especially in fruits since they produce the bluish or reddish coloration typical of the ripe fruits4. It is considered that fruits with intense bluish colors possess greater nutritional, medicinal, and antioxidants properties5, which have been associated with reducing coronary, neurological, and cardiovascular diseases6, cancer, diabetes, and inflammatory processes; this has been observed in various in vitro and in vivo7,8 studies.
The metabolic synthesis of flavonoids is well described for some species; however, its synthesis and anthocyanin concentration in Vaccinium floribundum are still unknown concerning the ripening process. Several genes involved with the synthesis of anthocyanin from phenylalanine have been identified; three of these are chalcone synthase (CHS), which participates at the beginning of the biosynthetic route, dihydroflavonol 4-reductase (DFR) and anthocyanin synthase (ANS) that are practically observed at the end of the synthesis4,9.
Molecular analyses are tools for managing biodiversity for the conservation and monitoring of endemic species and natural populations9. One of the molecular techniques used to observe gene expression patterns is the RT-QPCR (reverse transcriptase quantitative PCR)10, through which the concentration of one or more specific transcripts can be quantified11.
The objective of this research is to quantify the expression of the chalcone synthase, dihydroflavonol 4-reductase, and anthocyanin synthase genes that are directly involved in the biosynthesis of anthocyanin in two populations of Pichincha, namely Machachi and Pintag, since as mentioned by Li et al.12; there are insufficient specific genomic and transcriptomic profiles data to understand the molecular mechanisms associated with antioxidants, especially in Vaccinium species.
MATERIALS AND METHODS
Location and collection of samples
Ripe fruits of V. floribundum were collected in two wild populations of Pichincha, namely Machachi at Mejía with geographical coordinates 0°31′04.8″S 78°37′07.4″W and 3200 masl, and Pintag in Quito with geographical coordinates 0°24′00.0″S 78°24′00.0″W and 3000 masl.
According to the regulation for food collection of the INEN (Instituto Ecuatoriano de Normalización), 1kg of ripe fruits was collected from each native population and placed in a small tank with liquid nitrogen. The collected sample of V. floribundum, consisted of ripe fruits with the following characteristics: dark color in the peel, a diameter of 1 cm, and shrubs taller than 1.2 m. Although these moors are highly intervened, native populations of the species under study can be found.
RNA extraction
The extractions by columns were made using the Pure Link® RNA Mini Kit (Ambion, Life), according to the manufacturer's instructions. The final column was placed in a recovery tube, adding 50 µL of RNase free water to perform the elution and centrifuging it for 2 min at 12000 x g; afterward, 1 µL of RNase inhibition Protector (Roche) was added, and this preparation was retained at -20 ºC to be processed on the same day.
Reverse transcription
A reverse transcription kit was used to obtain the cDNA, specifically a transcriber First Strand cDNA synthesis kit and Random Primer of Roche, used according to the manufacturer's directions. The final microtube was taken to the conventional ThermoCycler (Labnet) using a protocol of 10-min at 25 ºC, 30-min at 55 ºC, 5-min at 85 ºC, and cooling at ∞ 4 ºC; finally, the cDNA was obtained and quantified in the Fluorometer Qubit® 2.0.
Elaboration of the standard curve
Serial solutions of the cDNA were prepared, obtained from a ripe V. floribundum fruit collected in Machachi, and the glyceraldehyde-3-phosphate dehydrogenase gene was used as a control to establish a patterned curve since it is a housekeeping gene.
Principio del formulario
The samples were quantified using Qubit® 2.0 Fluorometer with the Qubit® DNA Assay Kit (Life), following the instructions of the manufacturer; this was carried out using the DsDNA High sensitivity program, obtaining specific concentrations that will be entered into the RT-qPCR equipment for expression analysis of the genes.
With the quantified cDNA, amplification was performed using the Light Cycler Fast Start DNA MasterPLUS SYBR Green I (Roche), with the primers of the glyceraldehyde-3-phosphate dehydrogenase gene to enter the data in the Light Cycler 2.0 software (Roche).
In this experiment, programming of absolute quantification was performed, where the data obtained from the concentrations are entered through the Qubit, which are the positive standard controls for the subsequent analysis of the interest samples13.
The amplification run is initiated, giving the standard curve of cDNA concentration of the control gene, comparing the subsequent samples for their quantification in ng/uL.
Finally, it should be considered that the error values and the curve efficiency must be from 0.0 to 0.3 points and from 1.8 to 2.2 points, respectively, to be further later used in the runs of the RT-qPCR for quantification of samples with unknown concentrations. The standard curves with the control gene are the basis for the quantification at RT-qPCR14.
Quantification in RT-qPCR
The Master Mix Light Cycler Fast Start DNA MasterPLUS SYBR Green I (Roche) was previously prepared. For the master mix of each sample, the protocol of 20 µL of the final volume was standardized with 9.8 µL of Ultra-Pure H2O, 0.6 µL of the first forward, 0.6 µL of the first reverse, in a 10 micromolar concentration, plus 4 µL of the Master Mix and 5 µL of the previously obtained CP V. floribundum cDNA. On the other hand, a negative control, H2O PCR grade, and positive control, a sample of Mortiño, were required for each of the genes, and two more capillaries were prepared15. The sequences of the first GAPDH used were F:5’CAAACTGTCTTGCCCCACTT3’, R:5’CAGGCAACACCTTACCAACA3’, for DFR, F:5’GAAGTGATCAAGCCGACGAT3’, R:5’ATCCAAGTCGCTCCAGTTGT3’, for CHS, F:5’CCAAGGCCATCAAGGAATG 3’, R:5’TGATACATCATGAGTCGCTTC3’, for ANS, F:5’TCTTCTACGAGGGCAAATGG3’, R: 5’ACAGCCCATGAAATCCTGAC3’3. In the case of the negative control, 5 µL of the sample were replaced by 5 µL of Ultra-Pure H2O, and for the positive control the capillaries were carefully centrifuged at 12000 x g for 15s to ensure that the specimen is at the bottom of the sample and have a correct reading.
The amplification protocol was programmed in the Light cycler 4.0 software (Roche): initial denaturation at 95 °C, for 1 cycle, then 60 denaturation cycles at 95 °C, hybridization at 60 °C and extension at 72 °C, to finally reach the last step of cooling for 1 cycle at a temperature of 40 °C. Finally, the previously saved external standard curve must be imported to obtain the quantitative data16.
4x2 Factorial statistical analysis
A factorial evaluation of the four genes was carried out in the INFOSTAT program 2018α, namely chalcone synthase, dihydroflavonol 4-reductase, anthocyanin synthase, and glyceraldehyde-3-phosphate dehydrogenase in the two populations from Pichincha; i.e., with this analysis, it can be reported the relationship of each of the independent variables: each of the genes and all the genes with each one population and with two populations.
Using the variance analysis (ANOVA) with α= 0.01, it is verified if two or more averages of two or more data groups are similar or if some differ significantly from the others17.
The Duncan test was also performed to compare the degree of similarity in the expression of the 4 genes involved in the biosynthesis of the anthocyanin and the relationship between the two populations since this statistical test is used to examine all the differences among means considering that the number of repetitions is constant.
RESULTS
Standard curve using the glyceraldehyde-3phosphate dehydrogenase gene.
The standard curve generated was effective, as it yielded an efficiency of 2.121 points and an error of 0.202 points, which are in the acceptable ranges required by the ThermoCycler LightCycler 2.0; hence, it is concluded that the data obtained are reliable.
Figure 1 shows a curve in which the X axis represents the logarithmic concentration of the serial dilutions data that were obtained in the Qubit, whereas the Y axis represents the CP or crossing point in which the samples generated the amplification; the blue line is the trend line between the two axes, and it represents the normalization of the curve.
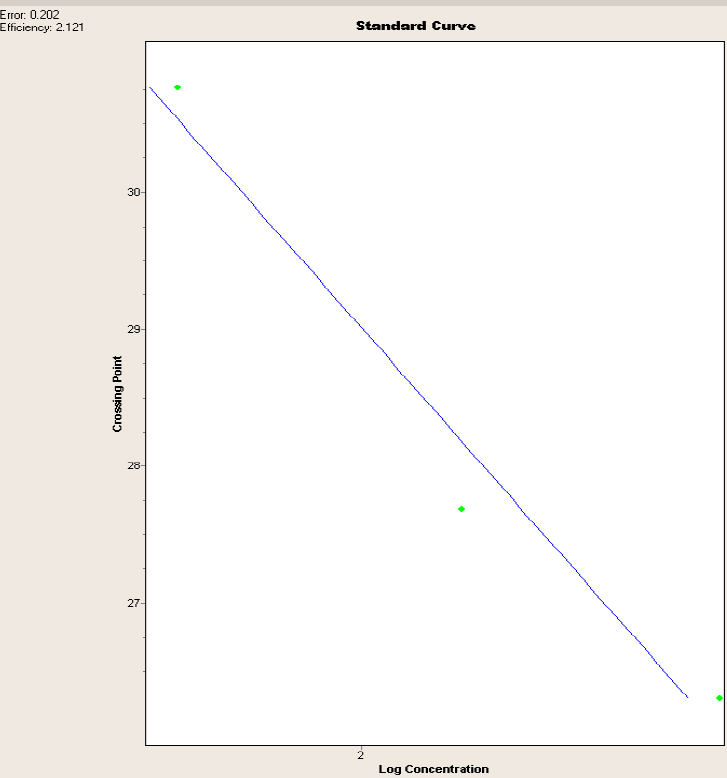
Figure 1. Standard curve for the V. floribundum with the glyceraldehyde-3-phosphate dehydrogenase gene. Error and efficiency are shown in the appropriate parameters.
Determination of the expression levels of the four genes in the species Vaccinium floribundum
The glyceraldehyde-3-phosphate dehydrogenase gene has the highest expression levels in both locations, with an average of 648.59 ng/ µL for Machachi (Table 1) and 667.32 ng/µL for Pintag (Table 2). Ten replications of the exact measurements were performed for each gene.
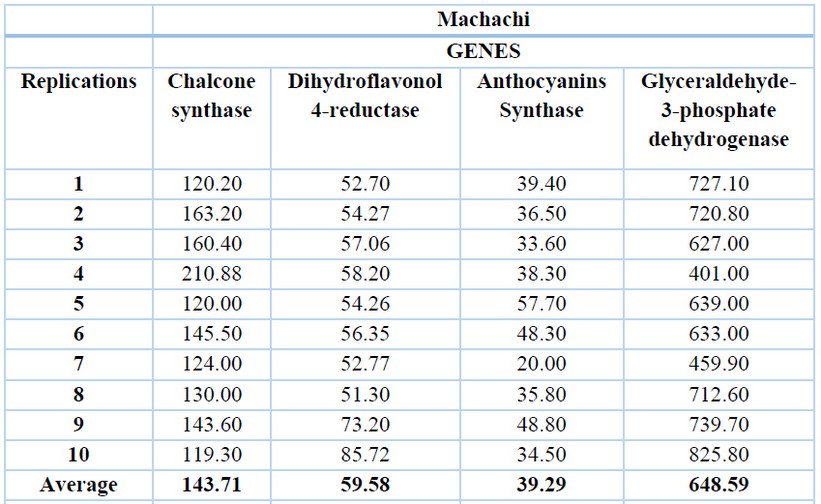
Table 1. cDNA concentrations (ng/µL) of the V. floribundum Machachi population for the four genes and their averages.
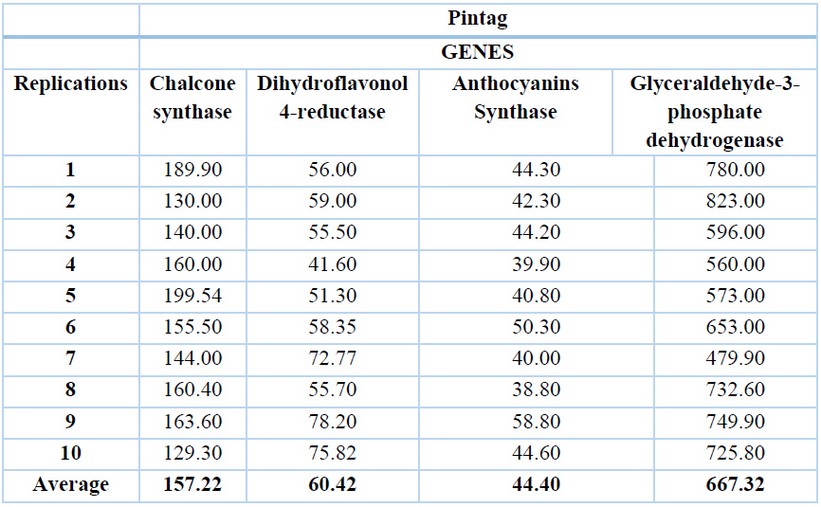
Table 2. cDNA concentrations (ng/µL) of the Pintag population of V. floribundum for the four genes and their averages.
The second gene with a higher expression level is the chalcone synthase with an average of 143.71 ng/µL in Machachi and 157.22 ng/µL in Pintag, then the dihydroflavonol 4-reductase gene with an average of 59.58 ng/µL in Machachi and 60.42 ng/µL for the population of Pintag; finally, the anthocyanidin synthase gene with an average of 39.29 ng/µL in Machachi and 44.40 in Pintag.
In Figure 2, one of the replications for the amplification curves of the four genes of the Machachi population is observed; this curve is inversely proportional to the concentration because while the curve rises in a minor cycle, the amount of the target will be higher. Four curves represent the four genes under study, plus the curve of negative control; the blue curve is almost linear until the end of the run without CP point. The green curve represents the glyceraldehyde-3-phosphate dehydrogenase gene, with a CP point of almost 27 cycles and a concentration of 727.10 ng/ µL; the red curve represents the chalcone synthase gene, with a CP of 38 cycles and a concentration of 120.20 ng/µL; the brown curve represents the dihydroflavonol 4-reductase gene, with a CP of 47 cycles and a concentration of 52.70 ng/µL; finally the black curve represents the anthocyanin synthase gene with a 60-cycle CP and a concentration of 39.40 ng/µL.
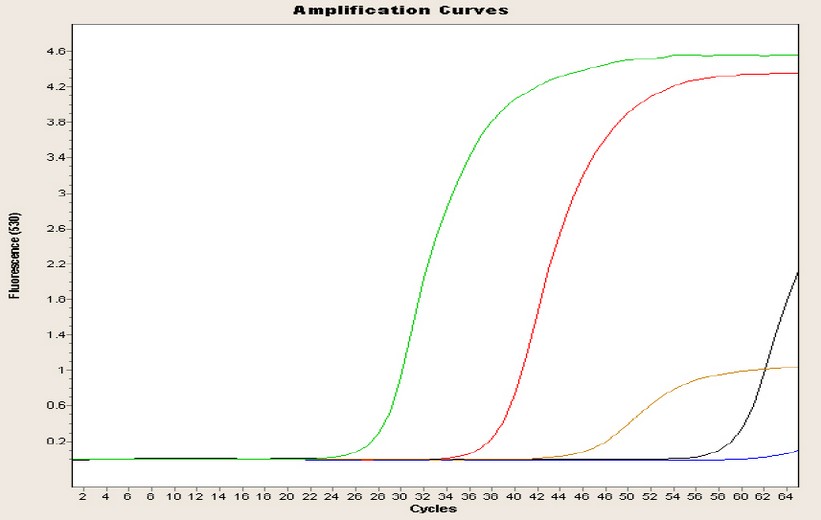
Figure 2. Amplification curve of the four population genes of Machachi population.
Regarding the Pintag population, Figure 3 shows one of the repetitions of the amplification curves of the four genes: the blue curve is almost linear until the end without CP point, and is the negative control. The green curve represents the glyceraldehyde-3-phosphate dehydrogenase gene with a 25-cycle CP point and a concentration of 780 ng/µL; the light blue curve represents the chalcone synthase gene with a 32-cycle CP and a concentration of 189.90 ng/µL; the brown curve represents the dihydroflavonol 4-reductase gene, with a 46-cycle CP and a concentration of 56 ng/µL; and finally, the black curve represents the anthocyanin synthase gene with a 59 cycle CP and a concentration of 44.30 ng/µL.
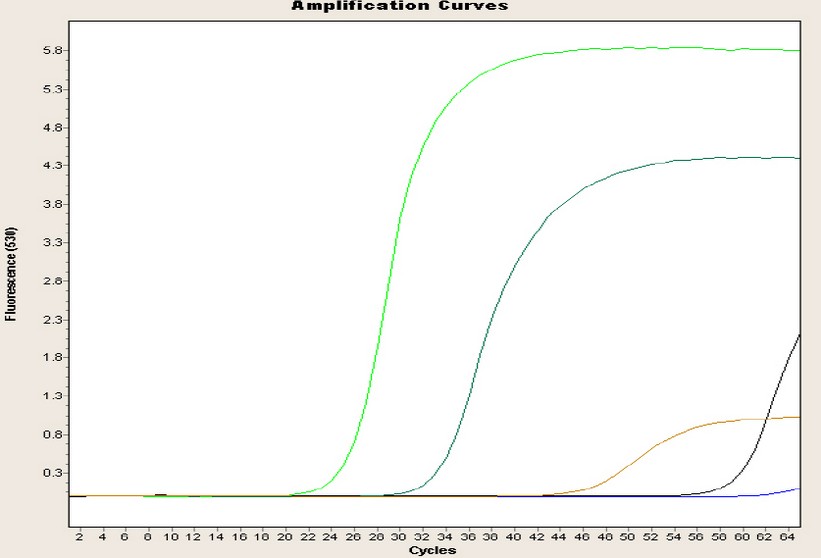
Figure 3. Amplification curves of the four genes of the Pintag population.
Comparison of the expression gene levels of Vaccinium floribundum in the two populations of Pichincha.
When comparing the two populations under analysis, no significant differences were found in any of the four genes (F = 0.47, p = 0.4965). On the contrary, the expression averages of each gene are very similar in both populations. In contrast, highly significant differences were observed when comparing the expression levels among the four genes analyzed (F = 433.24; p < 0.0001).
This study showed an expression level related to the biosynthetic route, where CHS has the highest expression level and ANS the lowest one. The differential expression of each gene observed in ripe fruits would be related to the maturation stage of this species and the function of each gene in the synthesis of anthocyanin. It would be essential to observe the expression of these genes in V. floribundum under different maturation stages. The results of this study (Duncan test) Show (Figure 4) that there are no significant differences in the expression levels when comparing the Machachi and Pintag populations. However, three expression levels can be observed; the anthocyanin synthase gene presented the lowest level, the intermediate level by the dihydroflavonol 4-reductase gene, and the highest expression level was observed in the chalcone synthase and glyceraldehyde-3-phosphate dehydrogenase (control) genes.
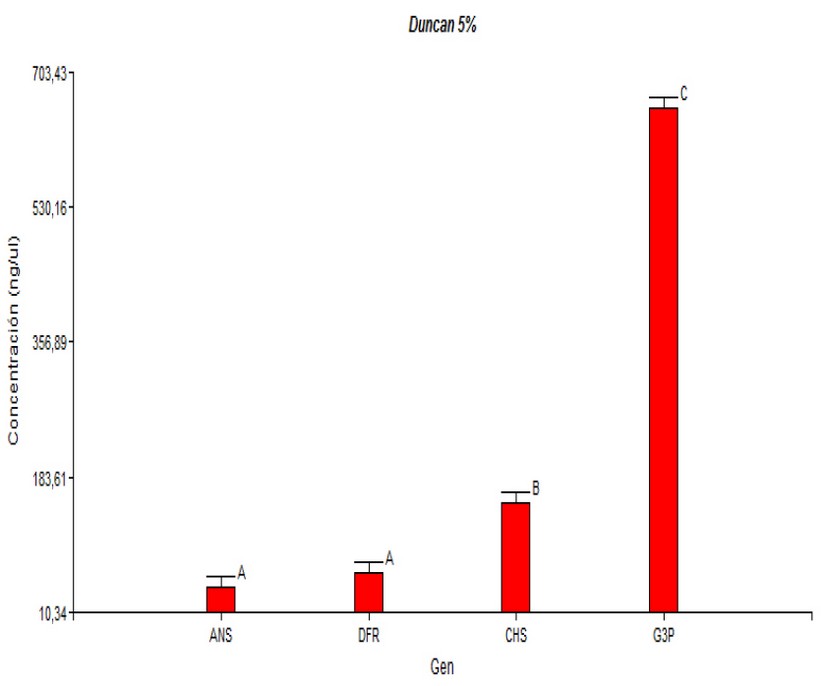
Figure 4. Duncan Test for comparing the averages according to the cDNA gene concentrations in ng/µL.
The ANOVA analysis among populations of V. floribundum (F = 0.08 and p = 0.9688) indicates no significant differences between such populations. Instead, it is observed that each of the genes analyzed (chalcone synthase, dihydroflavonol 4-reductase, anthocyanin synthase and glyceraldehyde-3-phosphate dehydrogenase) has a different expression level; this expression level is similar in both populations of V. floribundum; Chen et al.18 and Wang et al.10 showed that internal factors such as the maturation degree and environmental factors such as humidity, temperature and amount of light can affect the synthesis levels of anthocyanin; therefore, knowing the metabolic pathways can allow the implementation of management plans and avoid the extinction of this species19.
DISCUSSION AND CONCLUSIONS
The glyceraldehyde-3-phosphate dehydrogenase gene presents the highest concentrations concerning other genes, and therefore it enables the normalization of a standard curve and generates reliable data, as has been proven in other studies in Bilberry conducted by Jaakola et al.14 and in V. Myrtillus conducted by Martz et al.3. The results obtained indicate a high correlation in the expression of the chalcone synthase and dihydroflavonol 4-reductase found in Clivia miniata20, suggesting that these genes are subjected to coordinate the regulation of anthocyanin in the species, i.e., the two genes have direct action in the translation process regulating the final products. According to the expression values obtained for these two genes, something similar could be happening in the V. floribundum.
In the two populations, the anthocyanin synthase gene has the lowest expression level. It is possible that, as mentioned by Aguilera et al.21, some fruits belonging to the family of the Rosaceas, Vitaceae, Solanaceae, Ericaceae, and Passifloraceae are rich in anthocyanin, but in the case of Mortiño, the delphinidin is codified by another gene, the F3'5'H (Flavonoid 3'5'-'-hydroxylase), which is a crucial precursor compound that could be found in higher proportion compared to the anthocyanin synthase22.
There are no significant differences when comparing the expression level of the three genes (CHS, DFR, and ANS) among the populations since each gene has a similar expression level in both; therefore, the altitudinal differences do not affect the metabolic processes, and in this case the biosynthesis route of the anthocyanin in V. floribundum. These results agree with the research conducted by Li et al.23 and Wang et al.10, who observed the concentration of anthocyanin and flavonoids in Vaccinium uliginosum berries from seven locations in the Khingan Mountains only increased when there was a wide variation in altitude among populations.
The expression level of the CHS, DFR, and ANS genes is different and is related to the expression order in the biosynthesis path of anthocyanin, so the first to express and the one with the highest expression level is CHS, then DFR, and finally ANS. These data agree with Jaakola et al.14 in Bilberry, where procyanidins and quercetin were the main flavonoids in the early development stages of the berry, but their levels decreased dramatically during the maturation progress and the last stages. The content of anthocyanin increased significantly, and these were the main flavonoids in the ripe berry. Therefore, there is a correlation between the accumulation of anthocyanin and the expression of flavonoids during the ripening of berries.
Each of the genes presents a determining function in each stage of growth and maturation of the species, and it can be concluded that in the ripening stage when the fruit has a dark violet coloration and a diameter of about 1 cm, the gene that is most expressed is the chalcone synthase.
Many enzymes, structural genes, and regulatory elements have been observed as transcription factors involved in the biosynthesis of anthocyanin24. At the beginning of the biosynthesis, structural genes are expressed as chalcone synthase (CHS), chalcone isomerase (CHI), and flavanone 3-hydroxylase (F3H), which are necessary for the synthesis of anthocyanin precursors and derivatives of flavone and flavonols; then biosynthetic genes such as the dihydroflavonol 4-reductase (DFR) and the anthocyanin synthase (ANS) are expressed, which are necessary for the synthesis of anthocyanin and proanthocyanin22.
V. floribundum Kunth can be considered as a source of anthocyanin; therefore, the expression analysis of the genes involved in the biosynthesis route of anthocyanin in V. floribundum offers essential information that will enable the implementation of conservation programs for the species.
REFERENCES
1. Racines-Oliva M, Hidalgo-Verdezoto M, Vasquez-Castillo W. Domesticación de mortiño (Vaccinium floribundum Kunth.): frutal andino con gran potencial para la industria alimenticia. Agronomía Colombiana. 2016; 34:1-5.
2. Chiluisa-Utreras V. Vaccinium floribundum Kunth, reserva de antocianinas en los páramos. Tesis de postgrado en Biología de la Conservación de Pontificia Universidad Católica, Quito-Ecuador. 2015.
3. Martz F, Jaakola L, Julkunen-Tiitto R, Stark S. Phenolic Composition and Antioxidant Capacity of Bilberry (Vaccinium myrtillus) Leaves in Northern Europe Following Foliar Development and Along Environmental Gradients. J. Chem. Ecol. 2010; 36(9): 1017-1028. DOI: 10.1007/s10886-010-9836-9.
4. Jaakola L. New insights into the regulation of anthocyanin biosynthesis in fruits. Trends Plant Sci. 2013. DOI: 10.1016/j.tplants.2013.06.003.
5. Fernandes I, Faria A, Calhau C, de Freitas V, Mateus N. Bioavailability of anthocyanins and derivatives. Journal of functional foods. 2014; 7:54-66. DOI: 10.1016/J.JFF.2013.05.010.
6. Vasco C, Ruales J, Kamal-Eldin A. Total phenolic compounds and antioxidant capacities of major fruits from Ecuador. Food Chemistry. 2008; 111:816-823. DOI: 10.1016/j.foodchem.2008.04.054.
7. Norberto S, Silva S, Meireles M, Faria A, Pintado M, Calhau C. Blueberry anthocyanins in health promotion: A metabolic overview. Journal of Functional Foods. 2013; 5:1518-1528. DOI: 10.1016/j.jff.2013.08.015.
8. Kruger MJ, Davies N, Myburgh KH, Lecour S. Proanthocyanidins, anthocyanins and cardiovascular diseases. Food Research International. 2014; 59:41-52. DOI: 10.1016/j.foodres.2014.01.046.
9. Thomsen P, Willerslev E. Environmental DNA - An emerging tool in conservation for monitoring past and present biodiversity. Biological Conservation. 2015; 183:4-18. DOI: 10.1016/J.BIOCON.2014.11.019.
10. Wang Z, Jiang C, Wen Q, Wang N, Tao Y, Xu L. Deep sequencing of the Camellia chekiangoleosa transcriptome revealed candidate genes for anthocyanin biosynthesis. Gene. 2014; 538(1):1-7. DOI: 10.1016/j.gene.2014.01.035.
11. Efrose R, Ciobotari G, Morariu A, Pascu D, Branza M, Sfichi-Duke L. The use of qRT-PCR approach for comparative phenylpropanoid gene expression studies in two Rubus spp. 2010.
12. Li X, Sun H, Pei J, Dong Y, Wang F, Chen H, Sun Y, Wang N, Li H, Li Y. De novo sequencing and comparative analysis of the blueberry transcriptome to discover putative genes related to antioxidants. Gene. 2012; 511:54-61. DOI: 10.1016/j.gene.2012.09.021.
13. Huggett J, Dheda K, Bustin S, Zumla A. Real-time RT-PCR normalisation; strategies and considerations. Genes & Immunity. 2005; 6:279-284. DOI: 10.1038/sj.gene.6364190.
14. Jaakola L, Määttä K, Pirttilä AM, Törrönen R, Kärenlampi S, Hohtola A. Expression of Genes Involved in Anthocyanin Biosynthesis in Relation to Anthocyanin, Proanthocyanidin, and Flavonol Levels during Bilberry Fruit Development. American Society of Plant Biologists. 2002; 130(2):729-739. DOI: 10.1104/pp.006957.
15. Chiluisa-Utreras V, Medrano K, Cadena A, Acurio R. Quantification of FEND and ITUDI Anti-fungal Lipopetide Gene Expression in Bacillus megaterium using RT-qPCR. J Pure Appl Microbiol. 2020; 14(4): 2339-2349. DOI: 10.22207/JPAM.14.4.12.
16. Chiluisa-Utreras V, Vaca I, Chicaiza O, Peñaherrera S. Analysis of the expression of β-actin, RUANS and RUMYB10 genes involved in the biosynthesis of anthocyanin using RT-qPCR in Rubus niveus in the Rumiñahui Canton. Journal of Berry Research. 2017; 7(3):195-201. DOI: 10.3233/JBR-170156.
17. Sánchez-Otero, J. Introducción al Diseño Experimental. Impreso en Ecuador. 2013. ISBN: 9978-42-520-9.
18. Chen Q, Yu H, Tang H, Wang X. Identification and expression analysis of genes involved in anthocyanin and proanthocyanidin biosynthesis in the fruit of blackberry. Scientia Horticulturae. 2012;141:61-68. DOI: 10.1016/J.SCIENTA.2012.04.025.
19. Sexton, D. Gestión Social de los Recursos Naturales y territorios: Debates y propuesta pedagógica desde un contexto andino. CICDA. Quito; 2002.
20. Viljoen CD, Snyman MC, Spies J. Identification and expression analysis of chalcone synthase and dihydroflavonol 4-reductase in Clivia miniata. South African Journal of Botany. 2013; 87:18-21. DOI: 10.1016/J.SAJB.2013.03.010.
21. Aguilera-Otíz M, Reza-Vargas M del C, Chew-Madinaveita RG, Meza-Velázquez J. Propiedades funcionales de las antocianinas. Revista de Ciencias Biológicas y de la Salud. 2011;13(2):16-22. DOI: 10.18633/bt.v13i2.81.
22. Yuan Y, Ma X, Shi Y, Tang D. Isolation and expression analysis of six putative structural genes involved in anthocyanin biosynthesis in Tulipa fosteriana. Scientia Horticulturae. 2013; 153:93-102. DOI: 10.1016/j.scienta.2013.02.008.
23. Li R, Wang P, Guo Q, Wang Z. Anthocyanin composition and content of the Vaccinium uliginosum berry. Food Chemistry. 2011; 125:116-120. DOI: 10.1016/J.FOODCHEM.2010.08.046.
24. Rouholamin S, Zahedi B, Nazarian-Firouzabadi F, Saei A. Expression analysis of anthocyanin biosynthesis key regulatory genes involved in pomegranate (Punica granatum L.). Scientia Horticulturae. 2015; 186:84-88. DOI: 10.1016/J.SCIENTA.2015.02.017.
Received: 6 September 2021
Accepted: 10 October 2021
Viviana Chiluisa-Utreras1*, Doris Vela2, Ivonne Vaca1, Ramiro Acurio1, Javier Chicaiza3 y Sofía Peñaherrera3.
1 Research group BIOARN, Universidad Politécnica Salesiana, Quito - Ecuador.
2 Laboratory of Evolutive Genetics. School of Biological Sciences. Pontificia Universidad Católica, Quito, Ecuador.
3 Laboratory of Life Sciences, Universidad Politécnica Salesiana, Quito - Ecuador.
*Corresponding author: [email protected], Av. Isabel La Católica N. 23-52 and Madrid, Quito, Ecuador. Tel.: +593 23962800/Ext. 2110.
ORCID
Viviana Chiluisa-Utreras1 https://orcid.org/0000-0001-8647-1791
Doris Vela2 https://orcid.org/0000-0001-7690-7758
Ivonne Vaca1 https://orcid.org/0000-0002-4565-6623
Ramiro Acurio1 https://orcid.org/0000-0002-2305-4349.
