2023.08.01.44
Files > Volume 8 > Vol 8 No 1 2023
Long survival of Neisseria meningitidis in freeze-dried cultures maintained in potentially unsuitable conditions
1Centro de Investigaciones, Diagnóstico y Referencia. Instituto de Medicina Tropical Pedro Kourí (IPK), La Habana, Cuba.
2Departamento de Investigaciones. Escuela Latinoamericana de Medicina (ELAM), La Habana, Cuba.
*Corresponding author: Dr. Rafael Llanes. Autopista Novia del Mediodía km 6, entre Carretera Central y Autopista Nacional, Arroyo Arenas, La Lisa, La Habana, Cuba. Tel: 537 2553532, Fax 537 204 6051; email: [email protected]
Available from: http://dx.doi.org/10.21931/RB/2023.08.01.44
Dear Editor
Currently, diagnosis of Neisseria meningitidis in nasopharyngeal samples can be made by culture and nucleic acid amplification techniques.1 In Cuba, molecular diagnosis of meningococcal meningitis was introduced at the National Reference Laboratory for Neisseria (NRLN), in 2010, through a PCR that amplifies a fragment of the ctrA gene using the protocol described by Taha, 2000.2 This gene codes for a capsule protein that regulates the adhesion of N. meningitidis to the host, and 16 to 28% of meningococci isolates, especially in the nasopharynx, lack this gene.2 In contrast, the sodC gene, related to the production of superoxide-dismutase of this organism, is less sensitive to antigenic variation, hence its importance for molecular diagnosis in patients and asymptomatic carriers.3 Information on the meningococcal carriage is essential for public health policy,4. Still, the high number of invasive meningococci disease (IMD) affecting Cuba during the 1980s and the absence of molecular tools prevented its accurate microbiological diagnosis in carriers.5 This study aimed to identify
In Ciego de Avila province, N. meningitidis, by conventional and molecular methods in freeze-dried nasopharyngeal cultures of 50 Neisseria spp., recovered from carriers during 1987-1988, one of the most affected by a large epidemic of IMD in Cuba.5 Lyophilized material, which was preserved for 30 years, was reconstituted in 2 mL of nuclease-free sterile distilled water (Promega, USA). One milliliter was subcultured onto chocolate agar and incubated at 36.5-37°C for 18-24 hours, with 5-10% CO2, and the other milliliter was used to perform DNA extraction. Conventional methods used as sugar utilization and Vitek®2 automated system (bioMérieux, France) identified Neisseria species.QIAamp®DNA Mini and Blood Mini method, QIAGEN, Germany made 1 DNA extraction. Two PCR systems were used for molecular identification of meningococci, a simple PCR test that amplifies a 523 bp fragment of the ctrA gene,2 and a real-time PCR (rt-PCR) that amplifies a 127 bp fragment of the sodC gene.3 Serogroup identification of Neisseria meningitidis isolates was developed by slide agglutination using Remel™ Agglutinating Sera (Lenexa, USA). In addition, the main serogroups (A, B, C, W135, Y, X) of meningococci were investigated by rt-PCR,6 in positive samples identified by both simple and rt-PCR systems.
Pharyngeal carriage of N. meningitidis has been considered a prerequisite for the development of IMD and is known to be essential for transmission.4 In this study, ten isolates (20%) recovered from lyophilized material of nasopharyngeal carriers were identified as meningococci by culture, the standard gold method for detecting bacterial carriage.7 Nine of ten isolates were serogroup B, which was predominant during the epidemic of IMD in Cuba,5 and the other isolate was non-groupable.
Freeze-drying is more practical for the long-term preservation of
N. meningitidis cultures and optimal conditions for its conservation are refrigeration
(2-8°C) or freezing (-30 -70°C) temperatures, ampules are protected from the action of light and placed in an environment without humidity.8,9 These conditions produce good genetic stability of the product, with a longevity of up to 20 years.8 Moreira et al., in Cuba, obtained a 46.3% of survival of N. meningitidis lyophilized cultures after one year of storage at 4°C.10 Lyophilization is a very complex physical process affected by many parameters and variables such as growth medium, cell concentration, freezing rate, lyoprotectant, reconstitution medium, and time.11 In the current investigation, freeze-dried ampules were stored at room temperature and unprotected from light. However, its more prolonged survival of 30 years is noteworthy under these unsuitable conditions.8,9 Recently, Swain et al. demonstrated the relatively prolonged survival of the Cuban, New Zealand and Norwegian epidemic N. meningitidis serogroup B strains on inanimate surfaces for up to 8 days, depending on temperature and humidity, in comparison to other meningococci strains belonging to serogroup W135. In addition, carriage isolates appeared to survive better than invasive isolates, with a statistically significant difference (P = 0.002).12
Some authors recommend molecular tests for the identification and serogrouping of
N. meningitidis in cultures from carriers and lyophilized material.4,11 The end point-PCR results that amplify a fragment of ctrA gene detected N. meningitidis in 76% of cell suspensions of Neisseria spp. Investigated (Figure 1). Meanwhile, the rt-PCR that amplified a 127 bp fragment of the sodC gene identified meningococci in 100% of cell suspensions (Figure 2), which highlights the capacity of this rt-PCR system for the definitive identification of N. meningitidis. There are few studies on using sodC gene in nasopharyngeal cultures from carriers.3,4 Dolan et al. reported that sodC rt-PCR effectively identified 99.7% (624/626) of invasive and carriage N. meningitidis strains and was superior to the ctrA rt-PCR assay (71.6% 448/626).3 Jones et al. recently investigated three target genes (ctrA, sodC and porA) by whole-genome sequencing and rt-PCR. The ctrA gene was absent in a large percentage (58%, 54/93) of carriage isolates. However, both porA and sodC genes were well represented in the carriage collection (99% and 97%).13 In addition, Jordens & Heckels reported the development of a porA rt-PCR assay that identified several N. meningitidis isolates from carriers that were missed by using only the ctrA gene.7
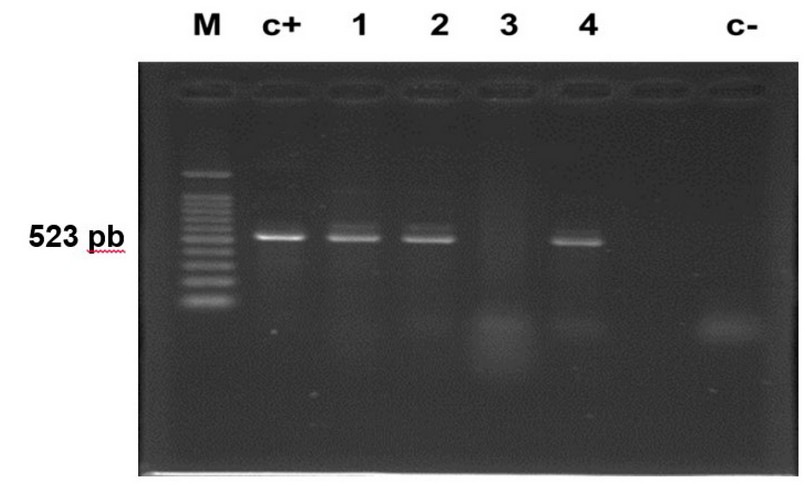
Figure 1. Amplifying Neisseria meningitidis ctrA gene in freeze-dried cultures of Neisseria spp. from carriers using a PCR with DNA extraction shown in 2% agarose gel stained with ethidium bromide, giving a 523 bp amplicon. Lanes: M: 100 bp DNA ladder; c+: positive control DNA of N. meningitidis ATCC 26195; lanes 1-4: DNA from carriers; c-: negative internal control (distilled water).
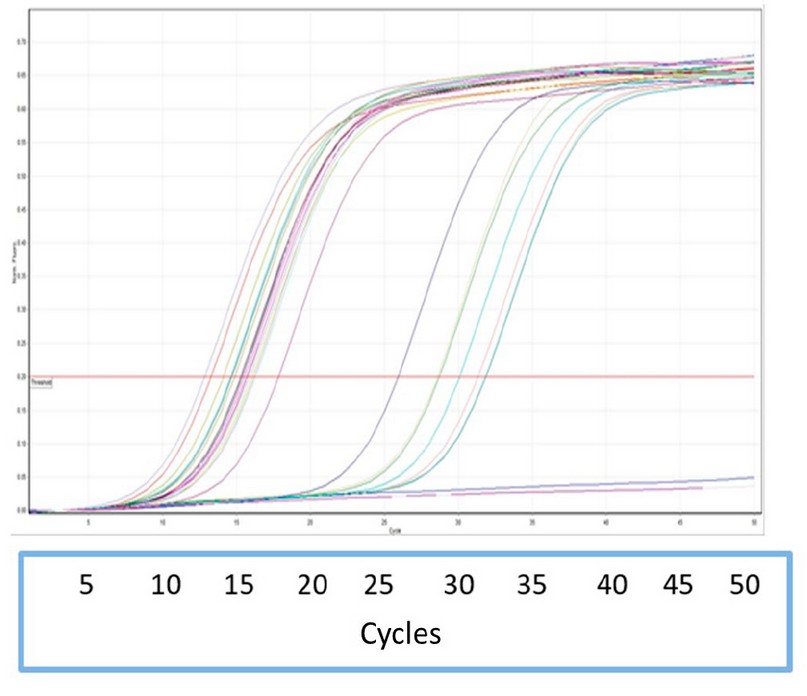
Figure 2. Specific amplification curves for real-time PCR with TaqMan that amplifies a 127 bp fragment of Neisseria meningitides sodC gene obtained from freeze-dried cultures of Neisseria spp in carriers. The horizontal line is the threshold.
In the current study, 68.4% (26/38) of cell suspension positive to N. meningitidis by both PCR belonged to serogroup B, 5.3% was serogroup C, and the remainder (26.3%) were non-groupable. Martínez et al., in a longitudinal study carried out on meningococci strains corresponding to nasopharyngeal carriers of the epidemic (1982-1992) and post-epidemic (1993-2002) stage in Cuba, detected a predominance of serogroup B (67.3%) in the epidemic phase and the non-agglutinable strains during the post-epidemic stage (70.7%).14 In Colombia, Moreno et al. also report a predominance of N. meningitidis serogroup B in carriers.15
The absence of comprehensive information for carriage in developing countries limits clarification of the epidemiology of IMD16. In the case of the Caribbean region, in particular, there is no previous report about the use of molecular methods for identifying or seron-grouping N. meningitidis in patients or carriers. The current study supports the usefulness of molecular tools in future studies of nasopharyngeal carriers in the Cuban population. In addition, this genetic material is helpful for further genomic characterization of Cuban meningococci strains by multi-locus sequence typing and/or other sequence methods. Genomic surveillance for N. meningitidis is fundamental for understanding pathogen evolution and disease epidemiology and can be significantly improved using culture-free methods.17
CONCLUSIONS
This study represents the first international report about a more prolonged survival of N. meningitidis in freeze-dried cultures from nasopharyngeal carriers that were kept under potentially unsuitable temperature, humidity and light exposure conditions. The sodC-based rt-PCR assay had an advantage over ctrA gene for detecting meningococci in cell suspensions of Neisseria spp. from lyophilized material obtained during the extensive epidemic of IMD in Cuba in the 1980s.
Acknowledgments
The authors acknowledged Dr. Yaxier de Armas and Prof. Imti Choonara for comments about the manuscript and Dr. Eldy Machado for English revision.
Conflict of interest: none.
REFERENCES
1. Rizek CF, Luiz AM, Assis GR, Figueiredo F, Levin AS, Lopes ME. Comparison of methods to identify Neisseria meningitidis in asymptomatic carriers. Rev Inst Med Trop Sao Paulo. 2016; 58: 60. doi: 10.1590/S1678-9946201658060. Last accessed [June 28 2022]
2. Taha MK. Simultaneous approach for nonculture PCR-based identification and serogroup prediction of Neisseria meningitidis. J Clin Microbiol. 2000; 38: 855-7. doi: 10.1128/JCM.38.2.855-857.2000. Last accessed [June 28 2022]
3. Dolan T, Hatcher CP, Satterfield DA, Jordan M, Bach MC, Linskot KB, et al. SodC-based real-time PCR for detection of Neisseria meningitidis. PloS one. 2011; 6: e19361. https://doi.org/10.1371/journal.pone.0019361. Last accessed [June 28 2022]
4. Coch CA, Silva de Lemos AP, Outeiro MC, Mendoza RA, Ballester T, Von Groll A, et al. Detection of Neisseria meningitidis in asymptomatic carriers in a university hospital from Brazil. Rev Argentina Microbiol. 2015; 47: 322-7. doi: 10.1016/j.ram.2015.08.004. Last accessed [June 28 2022]
5. Sierra-González VG. Vacuna cubana antimeningocócica VA-MENGOC-BC®: Treinta años de uso y potencialidades vigentes. VacciMonitor. 2020; 29(1):32-43. ISSN 1025-028X. Last accessed [27 June 2022].
6. World Health Organization. PCR for Detection and Characterization of Bacterial Meningitis Pathogens: Neisseria meningitidis, Haemophilus influenzae, and Streptococcus pneumoniae. Chapter 10. In: Laboratory Methods for the Diagnosis of Meningitis caused by Neisseria meningitidis, Streptococcus pneumoniae, and Haemophilus influenzae. 2nd ed. WHO/IVB.11.09, 2011. 323 pp.
7. Jordens JZ, Williams JN, Jones GR, Heckels JE. Detection of Meningococcal Carriage by Culture and PCR of Throat Swabs and Mouth Gargles. J Clin Microbiol. 2002; 40 (1): 75-9. doi: 10.1128/JCM.40.1.75-79.2002. Last accessed [June 27 2022]
8. Green LH. Culturing and Preserving Microorganisms. In: Goldman E & Green LH. Practical Handbook of Microbiology. 2015. Chapter 3. CRC Press, London. https://www.routledgehandbooks.com/doi/10.1201/b17871-5. Last accessed April 22 2022.
9. Del Puerto CA, Iglesias E, Morales T, Baños N, Nocedo MD, Carnota G, et al. Organización y manejo de la colección de cepas de referencia del Instituto Finlay. Vaccimonitor. 2009; 18: 20-4. ISSN 1025-028X. Last accessed [27 June 2022]
10. Moreira T, Iglesias E, Delgado H. Preservation of Neisseria meningitidis group B by freeze-drying. J Microbiol Meth. 1995; 23: 343-6.
11. Peiren J, Buyse J, De Vos P, Lang E, Clermont D, Hamon S, et al. Improving survival and storage stability of bacteria recalcitrant to freeze-drying: a coordinated study by European culture collections. Appl Microbiol Biotechnol 2015; 99: 3559–71. doi: 10.1007/s00253-015-6476-6 Last accessed [June 27 2022]
12. Swain CL, Martin DR, Sim D, Jordan TW, MacKishan JK. Survival of Neisseria meningitidis outside of the
host: environmental effects and differences among strains. Epidemiol. Infect. 2017; 145, 3525–3534. doi:10.1017/S0950268817002473. Last accessed [June 27 2022]
13. Jones CH, Mohamed N, Rojas E, Andrews L, Hoyos J, Hackins JC, et al. comparison of phenotypic approaches to capsule typing of Neisseria meningitidis by use of invasive and carriage isolate collections. J Clin Microbiol 2016; 45: 25-34. doi: 10.1128/JCM.01447-15
14. Martinez I. Neisseria meningitidis: contribución al transporte-conservación y caracterización de cepas aisladas en Cuba (1982-1992). Tesis presentada en opción al grado científico de doctor en ciencias médicas. Ciudad de La Habana, 2004. http://scielo.sld.cu › pdf › mtr05206. Last accessed [27 June 2022]
15. Moreno J, Sanabria O, Saavedra S, Rodríguez K, Duarte C. Caracterización fenotípica y genotípica de Neisseria meningitidis serogrupo B aisladas en Cartagena, Colombia, 2012-2014. Biomédica. 2014; 35(1): 138-43. doi: http://dx.doi.org/10.7705/biomedica.v35i1.2414. Last accessed [27 June 2022]
16. Serra L, Presa J, Christensen H, Trotter C. Carriage of Neisseria meningitidis in low and middle income countries of the Americas and Asia: A Review of the Literature. Infect Dis Ther. 2020; 9(2): 209–40. doi: 10.1007/s40121-020-00291-9. Last accessed [June 27 2022]
17. Itsko M, Topaz N, Ousmane-Traoré S, Popoola M, Ouedraogo R, Gamougam K, et al. Enhancing Meningococcal Genomic Surveillance in the Meningitis Belt Using High-Resolution Culture-Free Whole-Genome Sequencing. J Infect Dis 2022; 226: 729-37. doi: 10.1093/infdis/jiac1042022;226:729–37. Last accessed November 2, 2022
Received: December 23, 2022 / Accepted: January 30, 2023 / Published:15 February 2023
Citation: Gutierrez O; Martínez I; Feliciano O; Jerez L; Llanes R. Long survival of Neisseria meningitidis in freeze-dried cultures maintained in potentially unsuitable conditions. Revis Bionatura 2023;8 (1)44. http://dx.doi.org/10.21931/RB/2023.08.01.44
