2022.07.02.5
Files > Volume 7 > Vol 7 No 2 2022
Identification and phylogenetic characterization based on DNA sequences from RNA ribosomal genes of thermophilic microorganisms in a high elevation Andean tropical geothermal spring
Roque Rivas-Párraga1, Andrés Izquierdo1-6-7*, Karen Sánchez2, Darío Bolaños-Guerrón3,6, and Alonzo Alfaro-Núñez4-5
1 Life science and Agriculture Department. Universidad de las Fuerzas Armadas ESPE, Sangolquí, Ecuador;
2 School of Biological Sciences and Engineering, Yachay Tech University, San Miguel de Urcuqui, Ecuador
3 Department of Earth Science and Constructions, Geographical and Environmental Engineering, Universidad de las Fuerzas Armadas ESPE, Sangolquí, Ecuador
4 Clinical Biochemistry Department, Næstved Hospital, Ringstegade 57a, 4700 Næstved, Denmark
5 Section for Evolutionary Genomics, GLOBE Institute, University of Copenhagen, Øster Farimagsgade 5, 1353 Copenhagen K, Denmark
6 Centro de Nanociencia y Nanotecnología (CENCINAT), Universidad de las Fuerzas Armadas ESPE, Av. Gral. Rumiñahui s/n, P.O. Box 171-5-231B, Sangolquí, Ecuador Ecuador
7 Grupo de Investigación en Microbiología y Ambiente (GIMA), Universidad de las Fuerzas Armadas ESPE, Sangolquí, Ecuador
* Correspondence: [email protected]
Available from: http://dx.doi.org/10.21931/RB/2022.07.02.5
ABSTRACT
Several microorganisms can survive in harsh acid environments in geothermal springs at high temperatures across the Equatorial Andes Mountains. However, little is known about their physiological features and phylogenetic composition. Here we identify thermophilic microorganisms (bacteria, fungi, and microalgae) hosted in an almost unexplored geothermal spring (known as “Aguas Hediondas”). The phylogeny of the cultures was determined by analyzing physiological features and DNA sequences of PCR products for 16S rRNA, ITS, and 23S rRNA genes. Twenty pure cultures were isolated from the samples, including 17 for bacteria, one for cyanobacterium, one for eukaryotic microalgae, and one for fungus. Most bacterial strains were gram-positive, spore-forming, and bacilli (Bacillus). Cyanobacterium strain belonged to Chroococcidiopsis and the eukaryotic microalgae to Chlorophyta. The unique fungal strain isolated was closely related to T. duponti. Through our study, isolated thermophilic bacteria, microalgae and fungi from the “Aguas Hediondas” geothermal spring were characterized and identified. This study represents one of the first extensive molecular characterizations of extremophile microbes in the Tropical Equatorial Andes.
Keywords. microbial diversity; DNA markers; extremophiles; phylogenetics
INTRODUCTION
There has been a growing interest in studying extreme ecological niches and microorganisms living under extreme environmental conditions in the last decades. The increasing socio-economic and scientific relevance in discovering the biodiversity of extremophile microorganisms is primarily due to the direct and indirect (by-products) use in biotechnology and industries. Historically, a great diversity of microorganisms has been found in hostile conditions of geothermal environments.1–5 Microorganisms living in geothermal springs have developed several adaptations to survive hostile environments.6 For instance, these organisms can synthesize enzymes that work under high temperatures, high salt concentrations, high alkalinity or acidity conditions, and under high pressure.7,8 Their capacity to maintain their metabolic activity under these extreme conditions makes these enzymes (synthesized by thermophilic organisms) incredibly attractive for developing various human applications in diverse fields such as medicine, cosmetics, and the food industry.7,8 The culture-dependent method has played a crucial role in isolating microbes and preserving them for further research on biotechnological applications and developing new bioproducts.9
The biodiversity in geothermal springs is an adaptable response to the environmental conditions correlated with the mineral composition, pH, gases, salinity, redox potential, nutrients availability, and temperature variables.10,11 Additionally, climate change and anthropogenic activities can cause significant changes in the water chemistry of lentic and lotic ecosystems, which may affect the microbial biodiversity of these unique ecosystems. Biodiversity in tropical mountains is wealthy and high, particularly across the Andean mountains around Ecuador.12 Around 25 % of all terrestrial areas on Earth are mountain regions hosting more than 85% of the world’s species of amphibians, birds, and mammals, many exclusively restricted to mountains.13 Moreover, it has been well documented that hot spots in the tropical Andean mountains, including water reservoirs, can hold higher diversity than wet lowlands.14 The biodiversity of the different mountain ecosystems reflects the great importance of the evolutionary and ecological processes in these regions, a history worth understanding, preserving, and protecting.
Our study emphasizes microbial diversity in high-elevation Andean geothermal spring waters within these ecosystems. The survival of thermophiles depends on temperature regulation processes, activities, and behavior.15–17 It has been proposed that microorganisms shape ecosystems to make the environment more suitable for life.18–20 Evidence for symbiotic assemblages has been reported for diverse microbial communities in the microbe mats co-existing in hot springs.21,22 Main groups of microorganisms found in thermophilic environments, like archaea, bacteria, microalgae, and fungi, have adaptations to adverse conditions.15,23,24
Only a few studies have addressed the microbial and physicochemical characterization of “Aguas Hediondas” geothermal spring; however, they revealed only the diversity of microalgae communities without considering other important groups of microorganisms.25,26 Most existing reports have focused exclusively on tourism and geothermal energy production applications.27–30 It is believed by the local people that water from the geothermal spring has medicinal and healing properties.31,32 We find these proposed properties interesting to investigate, therefore relevant to characterizing the microbial communities in “Aguas Hediondas.”
This study aimed to identify thermophilic microorganisms in a little-explored hot spring in the Equatorial Andes using culture-dependent methods and classification based on DNA sequences from RNA ribosomal genes.
MATERIALS AND METHODS
Site characteristics and sample collection
Sediment and water samples were collected from the “Aguas Hediondas” geothermal spring located in Carchi province in northern Ecuador (00º 48.587’N & 77º 54.362’W) at 3428 m.a.s.l. The samples were taken from three different places in the same geothermal spring in April 2016. Three replicates were taken at each sampling point. Each replicate consisted of 50 ml of water and a portion of sediment from the bottom of the spring. The samples were maintained at 50 ºC during transportation and storage until the culture using a portable incubator (BIOBASE). The average temperature recorded in the three geothermal water points was 54.67±1.63 ºC. Environmental water and sediment samples were collected under permit # MAE-DNB-CM-2017-0071 granted by the Ministry of Environment of Ecuador. Water and sediment samples were sent for physicochemical analysis to a commercial service provider.
Media and culture conditions
Samples were cultured within six hours after collection. Sub-samples of 100 μl were spread on specific media for each group of microorganisms. The pour plate culturing method was used to get pure isolations. Bacteria were cultured on agar M9 (Difco) between 51 and 56 ºC for 72 hours.33 Cultivation of fungi was performed on PDA (Difco) supplemented with 50 mg l-1 of chloramphenicol (Sigma) at 55 ºC for 2 weeks.34 Microalgae were cultured on BG11 (Sigma) solidified with 1.5% Difco Bacto agar at room temperature (25 ºC) with illumination between 1000-2000 lux with 12 hours light and 12 hours dark photoperiod.35 Pure microalgae cultures were obtained by transferring part of each algal colony in 3 ml of liquid BG11 (Sigma). All the culture media were adjusted to the pH of the geothermal spring, around 5. Individual bacterial and fungal colonies were isolated by transferring to a fresh plate and incubated as indicated before prior to phenotypic characterization. Phenotypic features were examined according to colony pigmentation, texture, appearance, shape, and edge. After identification, bacterial and microalgal strains were cryopreserved in broth-glycerol (8:2) at -20ºC. Fungal isolates were preserved by culturing on PDA (Difco) dishes and then coated with mineral oil and stored at 4ºC.
Optical microscopy
All isolates were examined using a 100X magnification with a CX21 Olympus® microscope. Each isolate was characterized based on colony pigmentation, texture, appearance, shape, and edge (Supplemental Material). Mycelium structure was also observed in fungal isolates36. Bacterial isolates were Gram-stained for easier identification.
DNA extraction, PCR amplification, and DNA sequencing
Genomic DNA from bacteria, fungi, and microalgae was extracted by different protocols established by Moore et al.37, Weising et al.38, and Cai & Wolk,39, respectively. NanoDrop 8000 UV-Vis quantified DNA purity and concentration. Fragments of 16S rRNA, ITS, and 23S rRNA genes were amplified with specific primers (Table 1) through PCR. A 25 μl PCR reaction was carried out with the kit GoTaq® Green Master Mix from Promega. The PCR reaction mix was composed of 7 μl of ultrapure water, 12.5 μl of GoTaq® Green Master Mix (2X), 1.5 μl of each primer (10 μM) and 2.5 μl of DNA template. The protocol used for PCR amplification was 94°C for 5 min, 35 cycles of denaturation at 94°C for 30-sec primer annealing at 50°C for 1 min and extension at 72°C for 7 min and 30 sec with a final extension at 4°C for 10 min. The PCR products were visualized on 1 % agarose gels using GelStar™ dye. Amplified DNA fragments were sent for sequencing to a commercial sequencing service provider (Macrogen, South Korea).
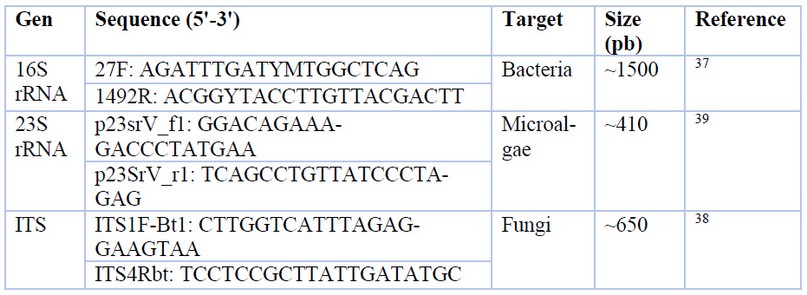

Table 1. PCR primer sequences used in this study.
Blast, sequence alignment, and phylogenetic analyses
DNA sequences used for this analysis are available under GenBank accession numbers MT765288-MT757927, MT757926, and MT764950-MT764966. To search for the similarity of the DNA sequences, the GenBank database (NCBI, National Centre for Biotechnology Information) was applied using the Basic Local Alignment Search Tool (BLAST) based on the most similar matches (>99% similarity). The obtained sequences were aligned using Geneious Prime® 2020.2.2 software through MUSCLE and ClustalW alignment methods and the online tool (https://mafft.cbrc.jp/alignment/server/index.html) for the MAFFT 7 alignment method. Phylogenetic reconstruction was performed with the Markov Chain Monte Carlo (MCMC) Bayesian approach implemented in BEAST version 1.10.4. Phylogenetic analysis was carried out with the GTR+Gamma as the best substitution model suggested by the software jModelTest for DNA sequences alignments.40 A non-parametric Bayesian Skyline (Piecewise-constant) coalescent model was used with a Strict molecular clock method. MCMC was developed with 20 million generations, subsampled every 1000 generations by applying a Random as the starting tree. Moreover, a Marginal Likelihood Estimation (MLE) using path sampling (PS) / stepping-stone sampling (SS), which performs an additional analysis after the standard MCMC chain has finished, was implemented. The MCMC/MLE analysis output was summarized using TreeAnnotator software included in the Beast package (.log files are provided in the supplemental material as a support of the analysis). The maximum Clade Credibility tree was produced after discarding 10 % of burn-in. The final tree was visualized through FigTree version 1.4.4.
RESULTS
The mean water pH measured was 4.8, while the sediment means pH was 5.8. Water and sediment samples showed distinct compounds and concentrations. Water contained 0.054 mg l-1 of arsenic, 83.1 mg l-1 of chlorides, 0.53 mg l-1 of iron, 2.5 mg l-1 of manganese 225 mg l-1 of sodium, 45.4 mg l-1 of potassium, 168.44 mg l-1 of magnesium, 72 mg l-1 of calcium, and 744.8 mg l-1 of sulphates. Water samples composition included volatile and non-volatile solids with 120 and 1 290 mg l-1 respectively. The sediment portion had the following compounds 1.3 mg kg-1 of cadmium, 29.3 mg kg-1 of copper, 312 mg kg-1 of potassium, 12 500 mg kg-1 of iron, 79 mg kg-1 manganese, and 1 860 mg kg-1 of magnesium. Sediments also contained organic matter, which represents approximately 60.59 % mean for all the samples. Electrical conductivity (EC), an indicator of water quality and inorganic constituents’ presence, was 0.1725 Sm-1 in water and 0.0164 Sm-1 in sediments. Higher EC levels denote higher total solids dissolved concentrations (TSD). EC is an indicator of dissolved minerals.
A total of 20 pure cultures were isolated, including 17 bacterial isolates, one cyanobacterium, one eukaryotic microalga, and one fungus. Most bacterial isolates were gram-positive spore-forming bacilli. The unique fungal strain isolated produced a grey dusty mycelium. Microalgae colonies were light green (eukaryotic) and bluish-green (cyanobacterium) (Figure 1).

Figure 1. Microbial strains isolated. (a) Bacteria (B. licheniformis). (b) Cyanobacterium (Chroococcidiopsis). (c) Eukaryotic microalgae (C. vulgaris). (d) Fungi (T. thermophilus).
Table 2 shows the relationships of isolated geothermal strains with the most similar sequences from the Genbank database.
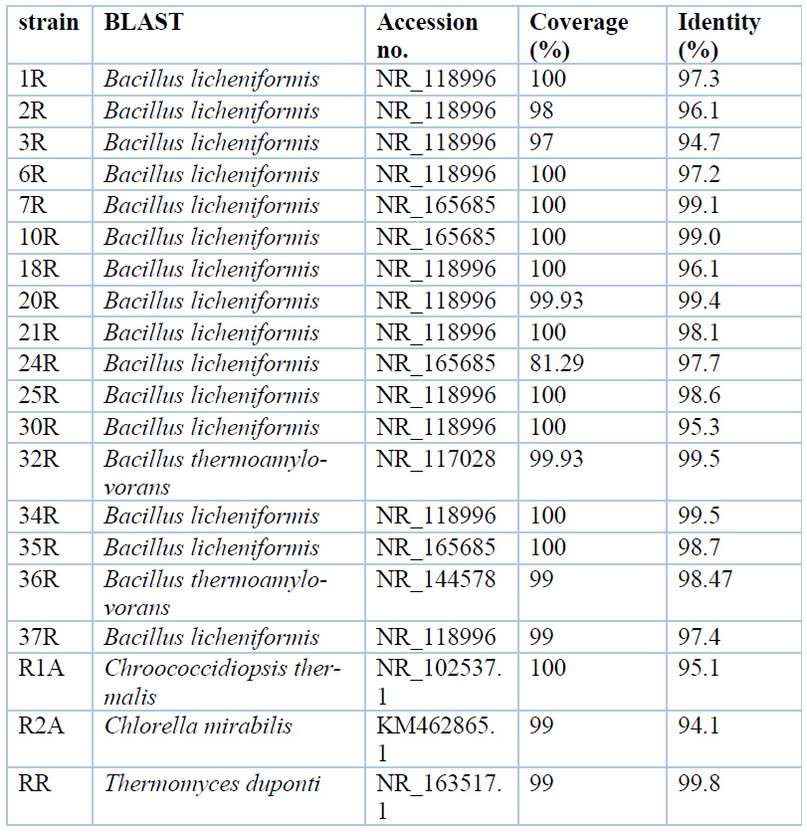
Table 2. Sequences from GenBank producing significant alignments
Figure 2 presents the phylogenetic tree of 17 isolated bacterial DNA strains. Two clades are marked, where the distance was compared by measures based on rRNA sequence divergence. All the bacteria (17) were contained within the genus Bacillus in the phylum Firmicutes. The nucleotide similarity in the ribosomal RNA gene sequences reveals that the bacterial strains were separated into two main groups: a branch closely related to Bacillus licheniformis (Figure 2a), and a group closely related to strains of Bacillus thermoamylovorans (Figure 2b).
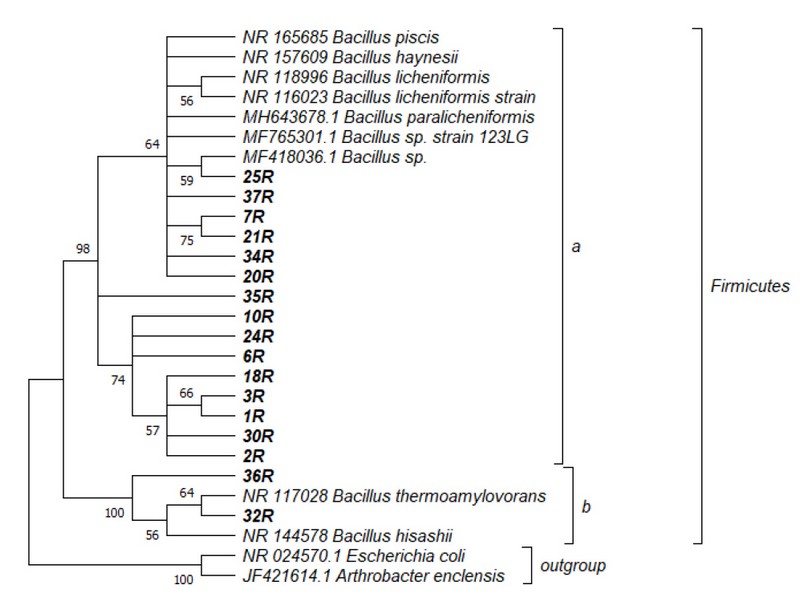
Figure 2. Bacteria phylogeny based on 16S rRNA gene analysis: isolated individuals closely related Bacillus strains. The phylogenetic tree was constructed using 1456 bp and maximum likelihood analyses under a GTR+Gamma Sites model. Numbers at nodes are support values generated from 1000 bootstrap replicates. GenBank accession no. MT764950-MT764966.
On the other hand, the BLASTn analysis exhibits that the only fungal isolate recovered from the hot spring was classified as closely related to Thermomyces duponti (Table 2).
Finally, the prokaryote microalgal isolate was a cyanobacterium classified as closely related to Chroococcidiopsis thermalis, and the eukaryote microalgal isolate was categorized as closely related to Chlorella mirabilis within Chlorophyta (Table 2).
DISCUSSION
The predominant presence of bacteria compared to fungi and microalgae may be due to the intrinsic geochemical properties of hot springs.41 However, the culture process must also be considered a restrictive issue for the determination of abundance. It is well established that approximately 1% of microorganisms can be readily cultivated in vitro.42,43 Because most microorganisms remain unculturable, the diversity in our study is inevitably underestimated using culture-dependent methods. For instance, M9, PDA, and BG11 are media that might not be the best for the recovery of ‘unculturable’ microorganisms from environmental samples.44 Methods, including the use of dilute nutrient media, could be a great approach to the recovery of ‘unculturable’ microorganisms adapted to oligotrophic conditions.45
Seventeen bacterial strains isolated in our study belong to the genus Bacillus, gram-positive and spore-forming. Similar findings were reported by Darland and collaborators,46 who observed that the genus Bacillus predominates in geothermal springs at Yellowstone National Park, especially those gram-positive with a pronounced tendency to form endospores. Bacterial strains isolated were classified as closely related to two species: B. licheniformis, and B. thermoamylovorans (Fig. 2). There are several reports of same species growing at similar conditions of temperature and pH in other geothermal springs around the world.47–54 B. licheniformis has been found in different geothermal springs in Ecuador, including Papallacta, Chachimbiro, Guapán, and Baños de Agua Santa.55–57
In the phylogenetic tree, E. coli and A. enclensis are located outside of the branch close to the genus Bacillus, as it should be expected, because these organisms belong to different phyla: Proteobacteria, Actinobacteria, and Firmicutes respectively.58–62 Wei Wang,63 also confirmed the position of B. thermoamylovorans concerning B. licheniformis. Furthermore, Yang Liu et al.64 and Christopher Dunlap et al.65 described that B. licheniformis and B. paralicheniformis are sister taxa sharing a common ancestor.
The Fungal ITS sequence analysis exhibited an association between T. duponti and the isolated strain (Table 2). T. duponti has been previously isolated from sediment samples in China's Tengchong Rehai National Park hot springs.66 Optimal growth conditions are similar with these found in “Aguas Hediondas” spring. Pan et al.66 established an optimal growth temperature for this fungus of 45-50°C and tolerable pH ranges of 4-12. Thermocyces dupontii belong to the order Eurotiales (Phylum Ascomycota).67,68
The examined samples in “Aguas Hediondas” geothermal spring contained prokaryotic and eukaryotic microalgaes. According to our analysis, isolated algae strains were classified as closely related to two species: Chroococcidiopsis thermalis, and C. mirabilis (Table 2). C. thermalis are part of the phylum Cyanobacteria.69 Moreover, C. mirabilis is an eukaryotic algae.69 Both species have been previously isolated from a similar environment. The genus Chroococcidiopsis has mainly been isolated from the interior and exterior of rocks in hot and cold deserts of the planet.66,70 It appears to be highly tolerant to desiccation.66,70 Other studies have identified strains of the genus Chroococcidiopsis in geothermal springs in northern Thailand with an optimal growth temperature of 50°C and pH 8.71 The eukaryotic microalgae C. mirabilis, which belongs to Chlorophyta, has been established as a phylum with an important role in algal communities from geothermal springs of Bulgaria.72 The data showed that 75 of the 200 species surveyed correspond to the phylum Chlorophyta, which can grow at temperatures in between 30 to 101ºC, and pH between 1-10.72 Our study is significantly different from local previously algal characterizations. Morales et al.25 reported the predominant presence of Closterium sp., Maougeotia sp., Navícula sp., Dictyosphaerium sp., and Ulothrix sp. in the Aguas Hediondas geothermal spring.
Ribosomal RNA barcodes were used for the identification and phylogenetic analysis of isolates because these kinds of markers are universal and composed, at the same time, of highly conserved as well as variable domains.73 Even if some researchers consider that ribosomal RNA is the best target for studying phylogenetic relationships, choosing the most suitable barcode could contribute to the resolution enhancement of the analysis presented in this study.74 Due to the increasing amount of information, 16S and 18S rRNA barcodes could improve the characterization of prokaryotic and eukaryotic microalgal isolates.
Because of their potential to produce thermostable extracellular enzymes with essential biotechnological uses, thermophilic and thermotolerant microorganisms are of significant economic value.75–77 The benefits of using thermostable enzymes for biotechnological processes at elevated temperatures include reduced risk of contamination by mesophilic microorganisms,78 decreased reaction-medium viscosity, increased bioavailability and solubility of organic compounds, and increased substrate and product diffusion coefficients, resulting in faster reaction rates.79 The species of microorganisms isolated in this study have been previously reported as producers of thermostable lipases,51 cellulases,80 xylanases,81 and pectinases.82 There are several reports of the use of these kinds of enzymes in biotechnological processes such as biobleaching of paper pulp,83 animal feed production,83 fermentation of sugars to obtain biofuel from cellulosic wastes,84,85 fruit juice extraction and clarification,84,85 refinement of vegetable fibers,86 degumming of natural fibers,86 curing of coffee,86 cocoa,86 and tobacco,86 and waste-water treatment.86 Further research is needed to elucidate the biotechnological applications of the isolated microorganisms in this study.
CONCLUSIONS
Through our study, isolated thermophilic bacteria, microalgae, and fungi from the “Aguas Hediondas” geothermal spring were characterized and identified. These results are confirmed by previous studies in the phylogeny and characterization of other geothermal waters. Our results represent an initial contribution to the study of thermophilic microorganisms in the geothermal spring “Aguas Hediondas”.
Author Contributions: Conceptualization, R.R.-P. and A.I.; methodology, R.R.-P., A.A.-N. and A.I.; software, R.R.-P. and A.A-N.; validation, R.R.-P.; formal analysis, R.R.-P., A.A.-N.; investigation, R.R.-P.; resources, A.I.; data curation, R.R.-P.; writing—original draft preparation, R.R.-P, K.S., D. B.-G.; writing—review and editing, R.R.-P, K.S., D. B.-G., A.A.-N., A.I.; visualization, R.R.-P.; supervision, A.I.; project administration, A.I.; funding acquisition, A.I. All authors have read and agreed to the published version of the manuscript.
Funding: This research was funded by Universidad de la Fuerzas Armadas-ESPE, grant number 2015-PIC-002.
Institutional Review Board Statement: Not applicable.
Informed Consent Statement: Not applicable.
Data Availability Statement: DNA sequences used for this analysis are available under GenBank accession numbers MT765288-MT757927, MT757926, and MT764950-MT764966.
Acknowledgments: Authors acknowledge the assistance of the Grupo de Investigación en Microbiología y Ambiente (GIMA) and Centro de Nanociencia y Nanotecnología (CENCINAT). Renato Naranjo for his collaboration in the research process. Thanks to SENESCYT-Ecuador for the economic support during this paper's writing process.
Conflicts of Interest: The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
REFERENCES
1. Kormas KA, Tamaki H, Hanada S, Kamagata Y. Apparent richness and community composition of Bacteria and Archaea in geothermal springs. Aquat Microb Ecol 2009; 57: 113–122.
2. Kublanov IV, Perevalova AA, Slobodkina GB, Lebedinsky AV, Bidzhieva SK, Kolganova TV et al. Biodiversity of thermophilic prokaryotes with hydrolytic activities in hot springs of Uzon Caldera, Kamchatka (Russia). Appl Environ Microbiol 2009; 75: 286–291.
3. Adams MWW. Biochemical diversity among sulfur-dependent, hyperthermophilic microorganisms. FEMS Microbiol Rev 1994; 15: 261–277.
4. Stetter K. Ultrathin mycelia-forming organisms from submarine volcanic areas having an optimum growth temperature of 105 °C. Nature 1982; 300: 258–260.
5. Stetter K. Diversity of extremely thermophilic archaebacteria. In: Brock TD (ed). The thermophiles: general, molecular and applied microbiology. John Wiley, 1986, pp 39–74.
6. Morozkina EV, Slutskaia ES, Fedorova TV, Tugaĭ TI, Golubeva LI, Koroleva OV. Extremophilic microorganisms: biochemical adaptation and biotechnological application (review). Applied Biochemistry and Microbiology 2010; 46: 5–20.
7. Adams MWW, Kelly RM. Enzymes From Microorganisms in Extreme Environments. Chemical & Engineering News 1995; 73: 32–42.
8. Vieille C, Zeikus GJ. Hyperthermophilic enzymes: sources, uses, and molecular mechanisms for thermostability. Microbiol Mol Biol Rev 2001; 65: 1–43.
9. Egamberdieva D, Birkeland N-K, Panosyan H, Li W-J. Extremophiles in Eurasian Ecosystems: Ecology, Diversity, and Applications. Springer, 2018.
10. Stetter K. Extremophiles and their adaptation to hot environments. FEBS Lett 1999; 452: 22–25.
11. Li F-L. Thermophilic Microorganisms. Caister Academic Press, 2015.
12. Cuesta F, Peralvo M, Merino-Viteri A, Bustamante M, Baquero F, Freile JF et al. Priority areas for biodiversity conservation in mainland Ecuador. Neotropical Biodiversity 2017; 3: 93–106.
13. Rahbek C, Borregaard MK, Colwell RK, Dalsgaard B, Holt BG, Morueta-Holme N et al. Humboldt’s enigma: What causes global patterns of mountain biodiversity? Science 2019; 365: 1108–1113.
14. Rahbek C, Borregaard MK, Antonelli A, Colwell RK, Holt BG, Nogues-Bravo D et al. Building mountain biodiversity: Geological and evolutionary processes. Science 2019; 365: 1114–1119.
15. Kristjansson JK. Thermophilic Bacteria. CRC Press, 1991.
16. da Costa MS, Duarte JMC, Williams RAD. Microbiology of extreme environments and its potential for biotechnology. Elsevier Applied Science, 1989.
17. Danson MJ, Hough DW, Lunt GG. The Archaebacteria: biochemistry and biotechnology. Portland Press, 1992.
18. Azam F. Microbial Control of Oceanic Carbon Flux: The Plot Thickens. Science 1998; 280: 694–696.
19. van der Heijden MGA, Bardgett RD, van Straalen NM. The unseen majority: soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems. Ecol Lett 2008; 11: 296–310.
20. Ishii S, Ikeda S, Minamisawa K, Senoo K. Nitrogen Cycling in Rice Paddy Environments: Past Achievements and Future Challenges. Microbes Environ 2009; 26: 282–292.
21. Rozanov AS, Bryanskaya AV, Ivanisenko TV, Malup TK, Peltek SE. Biodiversity of the microbial mat of the Garga hot spring. BMC Evol Biol 2017; 17: 254.
22. Mackenzie R, Pedrós-Alió C, Díez B. Bacterial composition of microbial mats in hot springs in Northern Patagonia: variations with seasons and temperature. Extremophiles 2013; 17: 123–136.
23. Patel A, Matsakas L, Rova U, Christakopoulos P. A perspective on biotechnological applications of thermophilic microalgae and cyanobacteria. Bioresour Technol 2019; 278: 424–434.
24. Ellis DH. Thermophilic Fungi Isolated from a Heated Aquatic Habitat. Mycologia 1980; 72: 1030–1033.
25. Morales E, Luna V, Navarro L, Santana V, Gordillo A, Arévalo A. Diversidad de microalgas y cianobacterias en muestras provenientes de diferentes provincias del Ecuador, destinadas a una colección de cultivos. Revista Ecuatoriana de Medicina y Ciencias Biológicas 2017; 34: 129–149.
26. Carrillo Y. Caracterización hidroquímica de las aguas termales del sur del Ecuador. 2019.
27. Montenegro F, Ortiz A. La Geotermia, Fuente de Energía Alternativa para el Departamento de Nariño: Potencialidad, Usos y Beneficios. Universidad Cooperativa de Colombia, 2015.
28. Haraldsson IG, Lloret A. Geothermal baths, swimming pools and spas: examples from Ecuador and Iceland. United Nations University, 2014.
29. Carrera DV, Guevara PV. Fuentes termales del Ecuador. Universidad de las Fuerzas Armadas ESPE, 2017.
30. Bacci ME, Mujica E. Turismo rural en el Ángel, Carchi, Ecuador: perspectivas y posibilidades. CONDESAN, 2001.
31. Montenegro F, Rosero C, Hernández I, Ortiz A. Empleo de la energía geotérmica en salud: usos terapéuticos de las fuentes termales del volcán Chiles en Nariño--Colombia. Ciência e Sustentabilidade 2016; 2: 183–207.
32. Enriquez YK. Diseño de una ruta eco termal y cultural para turismo de salud en la parroquia rural de Tufiño, provincia del Carchi, con la finalidad de promover el desarrollo de la comunidad. 2018.
33. Sambrook J, Fritsch E, Maniatis T. Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, 1989.
34. Rinaldi MG. Use of potato flakes agar in clinical mycology. J Clin Microbiol 1982; 15: 1159–1160.
35. Cadena M, Molina D, Carvajal A, Ontaneda D, Morales E. Bioprospección de macrocolonias de Nostoc sp. en los andes ecuatorianos. Acta Bot Venez 2013; 36: 287–308.
36. Harris JL. Safe, low-distortion tape touch method for fungal slide mounts. J Clin Microbiol 2000; 38: 4683–4684.
37. Moore E, Arnscheidt A, KrÜger A, StrÖmpl C, Mau M. Section 1 update: Simplified protocols for the preparation of genomic DNA from bacterial cultures. In: Akkrmans A, Van J, De Bruijn F (eds). Molecular Microbial Ecology Manual. Kluwer Academic Publishers, 2008, pp 1905–1919.
38. Weising K, Nybom H, Pfenninger M, Wolff K, Meyer W. DNA Fingerprinting in Plants and Fungi. CRC Press, 1994.
39. Cai YP, Wolk CP. Use of a conditionally lethal gene in Anabaena sp. strain PCC 7120 to select for double recombinants and to entrap insertion sequences. J Bacteriol 1990; 172: 3138–3145.
40. Darriba D, Taboada GL, Doallo R, Posada D. jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 2012; 9: 772.
41. López-López O, Cerdán ME, González-Siso MI. Hot spring metagenomics. Life 2013; 3: 308–320.
42. Hugenholtz P. Exploring prokaryotic diversity in the genomic era. Genome Biol 2002; 3: REVIEWS0003.
43. Amann RI, Ludwig W, Schleifer KH. Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol Rev 1995; 59: 143–169.
44. Köpke B, Wilms R, Engelen B, Cypionka H, Sass H. Microbial diversity in coastal subsurface sediments: a cultivation approach using various electron acceptors and substrate gradients. Appl Environ Microbiol 2005; 71: 7819–7830.
45. Vartoukian SR, Palmer RM, Wade WG. Strategies for culture of ‘unculturable’ bacteria. FEMS Microbiol Lett 2010; 309: 1–7.
46. Darland G, Brock TD. Bacillus acidocaldarius sp.nov., an Acidophilic Thermophilic Spore-forming Bacterium. Journal of General Microbiology 1971; 67: 9–15.
47. Lee CC, Kibblewhite-Accinelli RE, Smith MR, Wagschal K, Orts WJ, Wong DWS. Cloning of Bacillus licheniformis xylanase gene and characterization of recombinant enzyme. Curr Microbiol 2008; 57: 301–305.
48. Narayan VV, Hatha MA, Morgan HW, Rao D. Isolation and characterization of aerobic thermophilic bacteria from the savusavu hot springs in fiji. Microbes Environ 2008; 23: 350–352.
49. Derekova A, Mandeva R, Kambourova M. Phylogenetic diversity of thermophilic carbohydrate degrading bacilli from Bulgarian hot springs. World Journal of Microbiology and Biotechnology 2008; 24: 1697–1702.
50. Simanjuntak S, Semuel MY, Welerubun M, Naharia O. Molecular Barcoding Based 16S rRNA Gene of Thermophilic Bacteria from Vulcanic Sites, Linow Lake, Tomohon. Materials Science Forum 2019; 967: 83–92.
51. Deive FJ, Álvarez MS, Morán P, Sanromán MA, Longo MA. A process for extracellular thermostable lipase production by a novel Bacillus thermoamylovorans strain. Bioprocess Biosyst Eng 2012; 35: 931–941.
52. Khiyami MA. Thermo-aerobic bacteria from geothermal springs in Saudi Arabia. African Journal of Biotechnology 2012; 11. doi:10.5897/ajb11.3339.
53. Yohandini H, Julinar, Muharni. Isolation and Phylogenetic Analysis of Thermophile Community Within Tanjung Sakti Hot Spring, South Sumatera, Indonesia. HAYATI Journal of Biosciences 2015; 22: 143–148.
54. Gomri MA, Khaldi TEM, Kharroub K. Analysis of the diversity of aerobic, thermophilic endospore-forming bacteria in two Algerian hot springs using cultural and non-cultural methods. Annals of Microbiology 2018; 68: 915–929.
55. Arias JE, Maisincho J, Koch A, Flores F, Naranjo R, Izquierdo A. Isolation and molecular identification of bacterial community from Papallacta geothermal spring in Ecuador. J Biotechnol 2016; S: S19.
56. Moposita F. Identificación de bacterias, microalgas y hongos termófilos de la fuente geotermal ‘Chachimbiro’ mediante técnicas microbiológicas y moleculares. 2017.
57. Naranjo Y, Koch A. Identificación de bacterias, hongos y algas mesófilos y termófilos de las fuetnes geotermales: El Riñón-Provincia del Azuay y Guapán-Provincia del Cañar. 2017.https://repositorio.espe.edu.ec/bitstream/21000/13250/1/T-ESPE-057293.pdf (accessed 28 Mar2021).
58. Yuan M, Yu Y, Li H-R, Dong N, Zhang X-H. Phylogenetic Diversity and Biological Activity of Actinobacteria Isolated from the Chukchi Shelf Marine Sediments in the Arctic Ocean. Mar Drugs 2014; 12: 1281–1297.
59. Sutcliffe IC. A phylum level perspective on bacterial cell envelope architecture. Trends Microbiol 2010; 18: 464–470.
60. Haakensen M, Dobson CM, Deneer H, Ziola B. Real-time PCR detection of bacteria belonging to the Firmicutes Phylum. Int J Food Microbiol 2008; 125: 236–241.
61. de Hoon MJL, Makita Y, Nakai K, Miyano S. Prediction of Transcriptional Terminators in Bacillus subtilis and Related Species. PLoS Comput Biol 2005; 1: e25.
62. Dastager SG, Liu Q, Tang S-K, Krishnamurthi S, Lee J-C, Li W-J. Arthrobacter enclensis sp. nov., isolated from sediment sample. Arch Microbiol 2014; 196: 775–782.
63. Wei Wang MS. Phylogenetic relationships between Bacillus species and related genera inferred from 16s rDNA sequences. Braz J Microbiol 2009; 40: 505–521.
64. Liu Y, Lai Q, Du J, Shao Z. Bacillus zhangzhouensis sp. nov. and Bacillus australimaris sp. nov. Int J Syst Evol Microbiol 2016; 66: 1193–1199.
65. Dunlap CA, Kwon S-W, Rooney AP, Kim S-J. Bacillus paralicheniformis sp. nov., isolated from fermented soybean paste. Int J Syst Evol Microbiol 2015; 65: 3487–3492.
66. Pan W-Z, Huang X-W, Wei K-B, Zhang C-M, Yang D-M, Ding J-M et al. Diversity of thermophilic fungi in Tengchong Rehai National Park revealed by ITS nucleotide sequence analyses. J Microbiol 2010; 48: 146–152.
67. de Vries RP, Gelber IB. Aspergillus and Penicillium in the Post-genomic Era. Caister Academic Press, 2016.
68. Suh DY, Hyun MW, Kim SH, Seo ST, Kim KH. Filamentous Fungi Isolated from Platypus koryoensis, the Insect Vector of Oak Wilt Disease in Korea. Mycobiology 2011; 39: 313–316.
69. Guiry MD, Guiry GM. AlgaeBase. World-wide electronic publication, National University of Ireland, Galway. http://www.algaebase.org/ 2010.
70. Caiola MG, Billi D, Imre Friedmann E. Effect of desiccation on envelopes of the cyanobacterium Chroococcidiopsis sp. (Chroococcales). European Journal of Phycology 1996; 31: 97–105.
71. Hayashi NR, Peerapornpisal Y, Nishihara H, Ishii M, Igarashi Y, Kodama T. Isolation and cultivation of thermophilic cyanobacteria from hot springs of northern thailand. Journal of Fermentation and Bioengineering 1994; 78: 179–181.
72. Stoyneva MP. Survey on green algae of Bulgarian thermal springs. Biologia Bratislava 2003; 58: 563–574.
73. Woese CR. Bacterial evolution. Microbiol Rev 1987; 51: 221–271.
74. Fox GE, Woese CR, Pechman KR. Comparative Cataloging of 16S Ribosomal Ribonucleic Acid: Molecular Approach to Procaryotic Systematics. International Journal of Systematic and Evolutionary Microbiology. 1977; 27: 44–57.
75. Bhalla A, Bischoff KM, Uppugundla N, Balan V, Sani RK. Novel thermostable endo-xylanase cloned and expressed from bacterium Geobacillus sp. WSUCF1. Bioresour Technol 2014; 165: 314–318.
76. Hung K-S, Liu S-M, Tzou W-S, Lin F-P, Pan C-L, Fang T-Y et al. Characterization of a novel GH10 thermostable, halophilic xylanase from the marine bacterium Thermoanaerobacterium saccharolyticum NTOU1. Process Biochem 2011; 46: 1257–1263.
77. Qiao W, Tang S, Mi S, Jia X, Peng X, Han Y. Biochemical characterization of a novel thermostable GH11 xylanase with CBM6 domain from Caldicellulosiruptor kronotskyensis. J Mol Catal B Enzym 2014; 107: 8–16.
78. Haki GD, Rakshit SK. Developments in industrially important thermostable enzymes: a review. Bioresour Technol 2003; 89: 17–34.
79. Kumar S, Nussinov R. How do thermophilic proteins deal with heat? Cell Mol Life Sci 2001; 58: 1216–1233.
80. Acharya S, Chaudhary A. Optimization of fermentation conditions for cellulases production by Bacillus licheniformis MVS1 and Bacillus sp. MVS3 isolated from Indian hot spring. Braz Arch Biol Technol 2012; 55: 497–503.
81. Maalej I, Belhaj I, Masmoudi NF, Belghith H. Highly thermostable xylanase of the thermophilic fungus Talaromyces thermophilus: purification and characterization. Appl Biochem Biotechnol 2009; 158: 200–212.
82. Rehman HU, Aman A, Nawaz MA, Qader SAU. Characterization of pectin degrading polygalacturonase produced by Bacillus licheniformis KIBGE-IB21. Food Hydrocoll 2015; 43: 819–824.
83. Kulkarni N, Shendye A, Rao M. Molecular and biotechnological aspects of xylanases. FEMS Microbiol Rev 1999; 23: 411–456.
84. Ando S, Ishida H, Kosugi Y, Ishikawa K. Hyperthermostable endoglucanase from Pyrococcus horikoshii. Appl Environ Microbiol 2002; 68: 430–433.
85. Baffi MA, Tobal T, Lago JHG, Boscolo M, Gomes E, Da-Silva R. Wine aroma improvement using a β-glucosidase preparation from Aureobasidium pullulans. Appl Biochem Biotechnol 2013; 169: 493–501.
86. Jayani RS, Saxena S, Gupta R. Microbial pectinolytic enzymes: A review. Process Biochem 2005; 40: 2931–2944.
Received: 10 December 2021 / Accepted: 20 January 2022 / Published:15 May 2022
Citation: Rivas-Párraga, R.; Sánchez, K.; Bolaños-Guerrón, D.; Alfaro-Núñez, A.; Izquierdo, A. Identification and phylogenetic characterization based on DNA sequences from RNA ribosomal genes of thermophilic microorganisms in a high elevation Andean tropical geothermal spring. Revis Bionatura 2022;7(2) 5. http://dx.doi.org/10.21931/RB/2022.07.02.5
