2022.07.03.42
Files > Volume 7 > Vol 7 No 3 2022
1 University Of Baghdad / Institute of Genetic Engineering and Biotechnology for Postgraduate Studies, Iraq.
2 University Of Anbar/College Of Dentistry,Iraq.e-mail [email protected]
3University Of Anbar /College Of Pharmacy, Iraq.e-mail [email protected]
*correspondenc:e-mail [email protected]
Available from: http://dx.doi.org/10.21931/RB/2022.07.03.42
ABSTRACT
Hereditary and environmental variables have a role in the development of breast cancer. This study aimed to examine the links between genetic Variations in the GSTP1 gene and Predisposition to breast cancer in an Iraqi population. The research included 40 Iraqi female breast cancer patients and 20 healthy volunteers. GSTP1-1695 A/G gene polymorphisms were investigated using polymerase chain reaction in Real-time (RT-PCR). The results showed the GSTP1 frequency of the wild GG genotypes was showed significantly (P<0.01) higher in healthy women in comparison with Breast cancer women (GG, 80% vs. 32.5%, respectively; furthermore, heterozygous AG genotypes were significantly higher in Breast cancer women in comparison with healthy women 42.5% vs. 20%, respectively at (P<0.01). While the mutant AA genotype (25%) in patient women appeared significantly (P<0.01) higher compared to healthy women (0.0%). Finally, we discovered a connection between GSTP1 polymorphisms and a higher chance of developing breast cancer in an Iraqi female population sample.
Keywords: glutathione S-transferase1, breast cancer, polymorphism.
INTRODUCTION
Most cancers are attributable to both hereditary and environmental causes and have been confirmed by molecular epidemiological research. The most frequent type of cancer is breast cancer—widespread breast cancer globally and the most common cause of cancer-related mortality. Breast cancer, which accounts for 18% of all female cancers in developed and developing countries, is a significant public health concern in developed and developing countries.1, 2. The risk of cancer development has increased with the finding of genetic enzyme polymorphisms that participate in carcinogen metabolism 3, 4, 5. Carcinogen-metabolizing enzymes participate in numerous chemical agents, including xenobiotics and sex hormones, which are activated and deactivated in this way. 6,7. Step II metabolic enzymes belong to a superfamily. Multiple carcinogens' detoxification catalyzes glutathione for chemotherapeutic medicines, environmental pollutants, and carcinogenic and diverse xenobiological substances. GST variants that GST-mediated detoxification of carcinogenic compounds that have been intensively explored as possible breast cancer susceptibility genes is impaired. 8, 9, 10, 11. On chromosome 11 (11q13), GSTP1 genes are located 12. GSTP1 is mainly present in the heart, spleen, and lungs, and In breast tissue, GST is the most prevalent. 9,13,14. A single point mutation in the GSTP1 gene causes the replacement of Ile for Val at codon105, resulting in a change in enzyme function. 5,13. As a result, this research aimed to see how GSTP1 polymorphisms affected breast cancer susceptibility in an Iraqi population sample.
MATERIALS AND METHODS
The current research included 40 patients diagnosed with breast cancer at the clinic between March 2020 and July 2020 from Oncology Teaching Hospital in Medical City. Before beginning care, each patient provided pertinent details such as age, menopause, number of children, history of lactation and breast cancer family history. Twenty healthy female volunteers served as the control group. Written informed consent was given to each study participant.
Specimen collection. Following an overnight, Every subject enrolled in the study had 5 mL of blood drawn and collected in EDTA tubes for DNA extraction to determine GSTP1 genetic polymorphisms. Genomic DNA was extracted immediately according to the manufacturer's instructions using a standard DNA Extraction Kit (Dsbio, China). Using a NanoDrop ND-1000, the purity of DNA was determined by calculating the absorbance ratio at 260 and 280 nm (A260/A280), DNA concentration, and 260/280 ratios.
Detection of GSTP1 polymorphism.
Genotypes of GSTP1-1695 A/G polymorphisms were determined by RT-PCR qPCR software and were used in conjunction with the QIAGEN Real-time PCR System ((Rotor-Gene Q, Germany). (allelic discrimination). The following primers for GSTP1-1695 A/G were used(designed using NCBI): forward primer: 5-CAGGGCTCTATGGGAAGGAC-3 and reverse primer: 5- CCCTTTCTTTGTTCAGCCCC -3. Probe sequences for target GSTP1 gene. Dye Fam-BHQ: 5- ACATCTCCCTCATCTACACCAAC-3 and Dye Vic-BHQ: 5-ATACGTCTCCCTCATCTACACC-3 (Biosearch Technologies). Application program RT-PCR as follows: Hold 1 50 0C for 15min. Hold 2 94 0C for 15min.Denaturation 95for 5 sec, annealing 60 0C for 20 sec, extension 720C for 15 sec,(step one repeating for five cycles), then denaturation 95 0C for 5 sec., annealing 60 0C for 20 sec., finally extension72 0C for 15 sec.(step two Repeating for 40 cycles). The component used in genotype RT-PCR for GSTP1gene included forward primer 1μl, revers primer1μl, forward probe 1μl, reverse probe 1μl, probe master mix 10μl (WizPure™ qPCR Master (PROBE), South Korea, Dnase free water 3μl finally DNA3μl.The total volume was 20 μl. The SPSS software was used to examine the impact of various variables on study parameters. To compare percentages, the Chi-square test was utilized in this study.
RESULTS
According to the results achieved by PCR, The GSTP1 frequency of the wild GG genotypes was shown significantly (P<0.01) higher in healthy women in comparison with Breast cancer women (GG, 80% vs. 32.5%, respectively; furthermore, heterozygous AG genotypes were significantly higher in Breast cancer women in comparison with healthy women 42.5% vs. 20%, respectively at (P<0.01). While the mutant AA genotype (25%) in patient women has appeared significantly (P<0.01) higher compared to healthy women (0.0%), however, there were no significant differences in wild GG, heterozygous AG and mutant AA genotypes in females patient. According to genetic models, the frequency of dominant AA+AG genotype was significantly(P<0.01) in female patients (67.5%) than in control (20%). However, the frequency of recessive GG+AG genotypes was significantly (P<0.01) higher in healthy women (100%) compared to breast cancer women (75%), while mutant AA genotype was more in patient women than control (25% and 0.0%) respectively, the frequency of additive 2AA+AG genotypes were significantly(P<0.01) increased in breast cancer female (92.5%) compared to healthy female (20%) ( Figure 1 and Table 1).
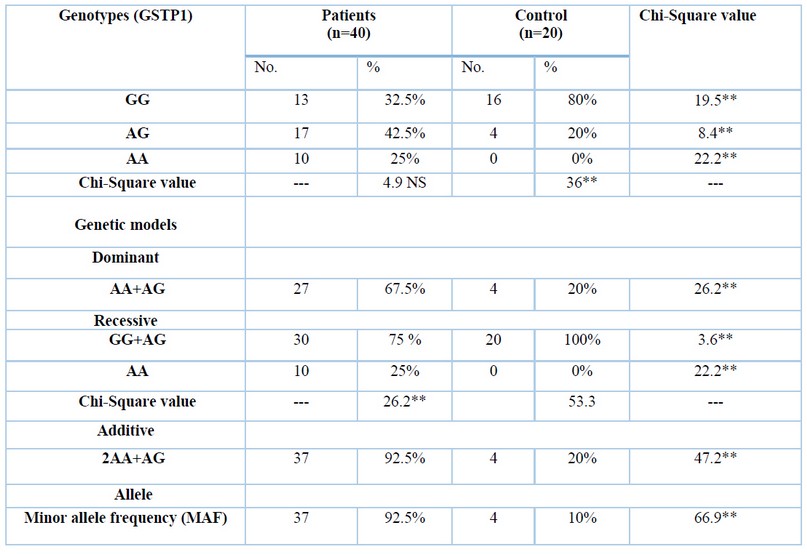
Table 1. Distribution of genotype and allele frequencies of GSTP1 gene in patients and controls.
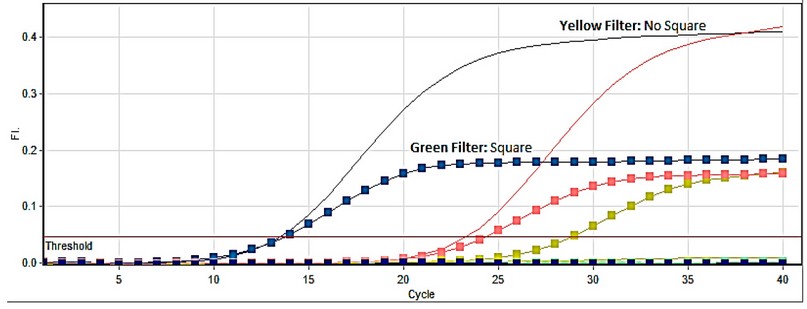
Figure 1. Determining GSTP1 gene polymorphism by RT-PCR using The Gentra Puregene Blood Kit ( Fam + Vic ) dyes.
DISCUSSION
Breast cancer (BC) is a condition affected by genetic and environmental factors. Twenty percent to twenty-five percent of all BC cases are caused by genes linked to susceptibility to the disease 15. GSTs, which are involved in the cell's detoxification process, contain a common gene (GSTP1) related to the progression of BC 16. In this research, we looked at the connection between GSTP1 genotypes With the risk of breast cancer in a group of Iraqi women. A substantial relationship between GSTP1 polymorphisms and the risk of breast cancer has been identified in the current investigation. Our findings agreed with Sergentanis and Economopoulos 17, who discovered that GSTP1 polymorphisms were correlated with a higher incidence of breast cancer in Chinese people but disagreed with many authors 8, 18, 19, 20, 21, 13, who observed that Genes were involved in the process of detoxification thought to be potential candidate genes for breast and another cancer susceptibility. GSTs are a group of enzymes involved in detoxifying a wide range of xenobiotics, including environmental carcinogens, hormones, and reactive oxygen species. Different liver functions and target tissue metabolizing enzymes may trigger individual differences in the metabolism of carcinogens. These differences could lead to varying vulnerabilities as breast cancer progresses. Increased oxidative stress and inflammation have long been linked to a high cancer risk 22. By conjugating glutathione, GSTs help to detoxify a variety of toxic xenobiotics. 23. The majority of GST substrates are electrophiles. Products of oxidative stress (i.e., oxygen species that are reactive.) (24) Cells that lack GSTP1 activity are more vulnerable to oxidative DNA damage. Tumors are prone to relevant genetic alterations during breast carcinogenesis as a result of the formation of DNA base adducts 14, 25. While some studies have looked into the connection between GST polymorphism and the risk of breast cancer, the link is still ambiguous and hotly debated.
CONCLUSIONS
This may be the first research to look at the relationship between GSTP1 polymorphisms and breast cancer risk in an Iraqi population. More research with larger samples is required to confirm our findings. Finally, This study discovered a connection between GSTP1 polymorphisms and an increased risk of breast cancer in an Iraqi female population.
Conflict of interest
No conflict of interest
Acknowledgment
We want to express our gratitude to all who support us.
Funds: Self
REFERENCES
1- Dević Pavlić, S., Ristić, S., Flego, V., Kapović, M., & Radojčić Badovinac, A.. Angiotensin-converting enzyme insertion/deletion gene polymorphism in lung cancer patients. Genetic testing and molecular biomarkers.2012; 16(7), 722-5.
2- Van Emburgh, B. O., Hu, J. J., Levine, E. A., Mosley, L. J., Case, L. D., Lin, H. Y. & Miller, M. S. Polymorphisms in drug metabolism genes, smoking, and p53 mutations in breast cancer. Molecular Carcinogenesis: Published in cooperation with the University of Texas MD Anderson Cancer Center.2008; 47(2), 88-99.
3. Mohammed,N.H.; Al-Taie, A. and Al-Sultany,F.H. Assessment of Predictive Confounders for the Prevalence of Breast Cancer among Iraqi Population:A Retrospective Study from Baghdad, Iraq . nternational Journal of Pharmacological and Pharmaceutical Sciences.2019;13(4).145-149.
4- Salinas-Sánchez, A. S., Sánchez-Sánchez, F., Donate-Moreno, M. J., Rubio-del-Campo, A., Serrano-Oviedo, L., Gimenez-Bachs, J. M., ... & Escribano, J. GSTT1, GSTM1, and CYP1B1 gene polymorphisms and susceptibility to sporadic renal cell cancer. In Urologic Oncology: Seminars and Original Investigations.2012;30(6)864-870.
5- Ramos, D. L., Gaspar, J. F., Pingarilho, M., Gil, O. M., Fernandes, A. S., Rueff, J., & Oliveira, N. G. Genotoxic effects of doxorubicin in cultured human lymphocytes with different glutathione S-transferase genotypes. Mutation Research/Genetic Toxicology and Environmental Mutagenesis.2011; 724(1-2), 28-34.
6- Kwon, D. D., Lee, J. W., Han, D. Y., Seo, I. Y., Park, S. C., Jeong, H. J. & Rim, J. S. Relationship between the glutathione-S-transferase P1, M1, and T1 genotypes and prostate cancer risk in Korean subjects. Korean Journal of Urology.2011; 52(4), 247.
7- Yeh, C. C., Lai, C. Y., Hsieh, L. L., Tang, R., Wu, F. Y., & Sung, F. C. Protein carbonyl levels, glutathione S-transferase polymorphisms and risk of colorectal cancer. Carcinogenesis.2010; 31(2), 228-233.
8- Ünlü, A., Ates, N. A., Tamer, L., & Ates, C. Relation of glutathione S‐transferase T1, M1 and P1 genotypes and breast cancer risk. Cell Biochemistry and Function: Cellular biochemistry and its modulation by active agents or disease.2008; 26(5), 643-647.
9- Šarmanová, J., Šůsová, S., Gut, I., Mrhalová, M., Kodet, R., Adámek, J., ... & Souček, P. Breast cancer: role of polymorphisms in biotransformation enzymes. European journal of human genetics.2004; 12(10), 848-854.
10- Reding, K. W., Weiss, N. S., Chen, C., Li, C. I., Carlson, C. S., Wilkerson, H. W., ... & Malone, K. E. Genetic polymorphisms in the catechol estrogen metabolism pathway and breast cancer risk. Cancer Epidemiology and Prevention Biomarkers.2009; 18(5), 1461-1467.
11- Oliveira, A. L., Rodrigues, F. F., Santos, R. E., Aoki, T., Rocha, M. N., Longui, C. A., & Melo, M. B. GSTT1, GSTM1, and GSTP1 polymorphisms and chemotherapy response in locally advanced breast cancer. Genet Mol Res.2010; 9(2), 1045-1053.
12- Mitrunen, K., Jourenkova, N., Kataja, V., Eskelinen, M., Kosma, V. M., Benhamou, S. & Hirvonen, A. Glutathione S-transferase M1, M3, P1, and T1 genetic polymorphisms and susceptibility to breast cancer. Cancer Epidemiology and Prevention Biomarkers.2001; 10(3), 229-236.
13- Vogl, F. D., Taioli, E., Maugard, C., Zheng, W., Pinto, L. F. R., Ambrosone, C. & Boffetta, P. Glutathione S-transferases M1, T1, and P1 and breast cancer: a pooled analysis. Cancer Epidemiology and Prevention Biomarkers.2004; 13(9), 1473-1479.
14- Henningson, M. C. K., Hietala, M., Bågeman, E., Olsson, H., & Jernström, H. Interactions between oral contraceptive status and GSTM1 and GSTT1 deletions on insulin-like growth factor-1 (IGF-1) plasma levels in young healthy women. Growth Hormone & IGF Research.2010; 20(6), 432-437.
15- Alshareeda, A. T., Negm, O. H., Albarakati, N., Green, A. R., Nolan, C., Sultana, R., ... & Rakha, E. A. Clinicopathological significance of KU70/KU80, a key DNA damage repair protein in breast cancer. Breast cancer research and treatment.2013; 139(2), 301-310.
16- Zheng, T., Holford, T. R., Zahm, S. H., Owens, P. H., Boyle, P., Zhang, Y., ... & Ali-Osman, F.. Glutathione S-transferase M1 and T1 genetic polymorphisms, alcohol consumption and breast cancer risk. British journal of cancer.2003; 88(1), 58-62.
17- Sergentanis, T. N., & Economopoulos, K. P. GSTT1 and GSTP1 polymorphisms and breast cancer risk: a meta-analysis. Breast cancer research and treatment.2010; 121(1), 195-202.
18- Gudmundsdottir, K., Tryggvadottir, L., & Eyfjord, J. E. GSTM1, GSTT1, and GSTP1 genotypes in relation to breast cancer risk and frequency of mutations in the p53 gene. Cancer Epidemiology and Prevention Biomarkers.2001; 10(11), 1169-1173.
19- Millikan, R., Pittman, G., Tse, C. K., Savitz, D. A., Newman, B., & Bell, D. Glutathione S-transferases M1, T1, and P1 and breast cancer. Cancer Epidemiology and Prevention Biomarkers.2000; 9(6), 567-573.
20- Cerne, J. Z., Pohar-Perme, M., Novakovic, S., Frkovic-Grazio, S., Stegel, V., & Gersak, K. Combined effect of CYP1B1, COMT, GSTP1, and MnSOD genotypes and risk of postmenopausal breast cancer. Journal of gynecologic oncology.2011; 22(2), 110-119.
21- Antognelli, C., Del Buono, C., Ludovini, V., Gori, S., Talesa, V. N., Crinò, L., ... & Rulli, A. CYP17, GSTP1, PON1 and GLO1 gene polymorphisms as risk factors for breast cancer: an Italian case-control study. BMC cancer.2009; 9(1), 1-14.
22- Ohshima, H., & Bartsch, H. Chronic infections and inflammatory processes as cancer risk factors: possible role of nitric oxide in carcinogenesis. Mutation Research/Fundamental and Molecular Mechanisms of Mutagenesis.1994; 305(2), 253-264.
23- Ketterer, B. Protective role of glutathione and glutathione transferases in mutagenesis and carcinogenesis. Mutation Research/Fundamental and molecular mechanisms of mutagenesis.1988; 202(2), 343-361.
24- Whalen, R., & Boyer, T. D. Human glutathione S-transferases. In Seminars in liver disease.1998;18(4) 345-358.
25- Cavalieri EL, Stack DE, Devanesan PD, Todorovic R, Dwivedy I, Higginbotham S, Johansson SL, Patil KD, Gross ML, Gooden JK, Ramanathan R. Molecular origin of cancer: catechol estrogen-3, 4-quinones as endogenous tumor initiators. Proceedings of the National Academy of Sciences. 1997;94(20):10937-42.
Received: 20 February 2022 / Accepted: 23 May 2022 / Published:15 August 2022
Citation: Rasheed M N, Hazim Hamoode R, Abdul-Jalil A A. Association of glutathione S-transferase 1 (GSTP1) polymorphisms with Breast Cancer susceptibility. Revis Bionatura 2022;7(3) 42. http://dx.doi.org/10.21931/RB/2022.07.03.42
