2022.07.01.29
Files > Volume 7 > Vol 7 No 1 2022
Epidemiological Molecular Analysis of Acinetobacter baumannii isolates using a multilocus sequencing typing and Global lineage
Heba A. Kadhom1* and Munim R. Ali2
1 Department of Biomedical Engineering, University of Technology, Baghdad, Iraq.
2 College of Science, Mustansiriyah University, Baghdad, Iraq.
*Correspondence: [email protected]
Available from: http://dx.doi.org/10.21931/RB/2022.07.01.29
ABSTRACT
The Multilocus sequence typing MLST method was used to recognize outbreaks of hospitals distinct clonal lineages of A. baumannii; these schemes appeared to provide largely concordant classifications that have been tools to evaluate the population structures of bacterial pathogens. One hundred fifty samples were collected from different specimens of patients within Baghdad hospitals (blood 40%, CSF 5%, urine 5%) between July 2019 to February 2020. Then identification all isolated as phenotypic detection and performed using PCR amplification of 16srRNA and blaOXA-51-like as genotypic detections. According to clinical and laboratory standards institute (CLSI) guidelines, Susceptibility testing was performed. Clonally analysis was performed by global lineage ICs correlated with multilocus sequence typing (MLST) when our data showed a very high rate of antimicrobial resistance in all hospital isolates, especially against colistin (8%) which determined the PDR isolates from other types also recorded 70% of isolates standing for carbapenems antibiotics (IMI 32%, MER70%& DOR 64%). Then already clustered into four groups according to multiplex PCR for two groups of three genes (ompA, csuE & blaOXA-51-like) where IC II was predominant in Iraq but in our strains founding ICI (38%) more prevalence one followed by IC0 (26%) then ICII and ICIII (20% &16% respectively). MLST used for detected the common sequence types (STs) of our selected 8 A. baumannii strains (IC0/A11, ICI/A6,48, ICII/A33,50,19 and ICIII/A1,36) were performed by using 7 housekeeping genes than were submitted in the MLST Pasteur scheme dataset (ID 5098, 5099, 5100, 5101, 5102, 5103, 5482 & 5483) followed by statistical eBurst analysis was done to study Clonal complexes (CCs). Identified 5 new STs (8, 444, 346, 1587 & 621) within Iraq and new one ST (1830) worldwide.
Keywords. Acinetobacter baumannii, antibiotic resistance, carbapenem, global lineage, MLST, eBURST, clonal complex, molecular typing, ST.
INTRODUCTION
The established standard for molecular typing method is multilocus sequence typing (MLST), which has been designed to study population structures of bacterial pathogens where two MLST schemes are both widely used (Oxford & Pasteur) 1. Both schemes produced largely concordant classifications with "true" phylogenic relationships 2. Briefly, MLST is based on the seven housekeeping genes sequenced analysis "gltA, gyrB, gdhB, recA, cpn60, gpi and rpoD". All these genes must be amplified then sequenced to submit within Pub MLST database, which led to finding the allelic number and the STs were assigned to each isolate with the seven allelic profiles 3. However, all datasets recorded supplementary analysis with different applications as in upon of the eBurst algorithm to describe the information of STs origins with multiple clonal complexes (CCs) and the investigation of disequilibrium linkage among alleles at the seven housekeeping genes 4.
For the classification of worldwide clones and produced accurate results was applying widely were used designed two multiplexes PCRs selectively amplify and sequence the ompA, csuE, and blaOXA-51-like genes that are under selective pressure and assign A. baumannii strains to different sequence groups (SG), which corresponding to international clonal lineages upon this can be identified using other approaches, making it a useful preliminary tool for studying local epidemiology 5, 6, 7. The rising use of monitoring of multidrug-resistant bacteria at the molecular level in the previous period of time constraints, so A. baumannii necessitates the search for reliable typing methods limitation in the time, labor, and costs. Global clone 1 (GC1) and global clone 2 (GC2) were two significant clones responsible for most of these outbreaks where referred to as international clones (IC) 1 and 2 8. Within each clonal complex, the latter procedure resulted in separate lineages or sub-lineages 9. The repeated replacement within genes further caused variations that delineate the capsule and lipooligosaccharide's outer core structure 10. Hence, the heterogeneity within each close relative producing outbreaks or spreading at the local, national, or global level can be distinguished using the clonal complex and variability 11. Finally, these more common methods are currently used to analyze the underlying genetic differences among isolates and consider a golden standard 12, 13. The MLST scheme provides a high level of resolution and an excellent tool for studying the population structure and long-term epidemiology of A. baumannii.
MATERIALS AND METHODS
Clinical samples collection
For 8 months of study (from August 2019 till March 2020), 50 A. baumannii were isolated from patients with different infection sites attending different hospitals in Baghdad\Iraq. The A. baumannii isolates were obtained from blood (40), urinary tract infections (5) and CSF (5). In a nutshell, samples were distributed on MacConkey agar plates and incubated at 37°C overnight. Petri plates were submerged with bacteria species in several samples, particularly UTI samples, making isolation challenging. As a result, only isolated colonies (one per sample) exhibiting unique A. baumannii morphology were chosen for subsequent analysis. Traditional biochemical tests (Gram staining, oxidase, indole, urease, citrate, and methyl-red-Voges-Proskauer) and the API 20E system (BioMérieux, La Balme Les Grottes, France) were used to identify the species. Finally, the A. baumannii isolates were preserved at –80°C in a 15 % (V/V) glycerol brain heart infusion.
Genotyping detection for isolates
Isolates were genetically confirmed according to the existence of genus and species- specific gene (16srRNA & oxa 51, respectively). Specific two primers, "240 bp & 353bp" employed and conditions' steps listed in table 1 & 2 respectively.
Antibiotic susceptibility test
The performance standard for antimicrobial disk susceptibility testing evaluated susceptibility using the agar disk diffusion technique 14. All of which were examined belonged to one of the most common antibiotic families being used to treat A. baumannii infections. The following antimicrobial agents have tested: amoxicillin (AMX), augmentin (AMC), piperacillin (PRL), Ticarcillin/ Clavulanic acid (TIM), Ampicillin/ sulbactam (SAM), Ticarcillin (TICER), Ceftriaxone (CRO), Cefotaxime (CTX), Ceftazidime (CAZ), Meropenem (MEM), Doripenem (DOR), Imipenem (IMI), Gentamicin (GEN), Amikacin (AK) Tobramycin (TOB), Ciprofloxacin (CIP), Levofloxacin (LEV), Gatifloxacin (GATI), Tetracycline (TE), Tigecycline (TG), Doxycycline (DXT), sulfa drug (SXT), Polymyxin B (PB) finally Colisten (COL).
Identification of housekeeping genes
To retain genetic diversity during storage, isolates were kept at -70°C in 20 % (vol/vol) glycerol in LB medium and cultivated overnight on MacConkey agar at 37°C. A colony's loopful was suspended in 500 microliters of distilled water. A QIAquick PCR Purification Kit was used to recover bacterial DNA (Qiagen, USA). DNA was kept at -20°C until it was needed.
Epidemiological typing
Global lineage (GL)
Three-locus dual assay multiplex PCR (M-PCR) was used to detect the international clone (IC) lineages of A. baumannii isolates, which selectively amplified the outer membrane protein A (ompA), chaperone–subunit usher E (csuE), and intrinsic carbapenemase (blaOXA-51-like) genes. Only the ompA fragment was amplified in the IC II M-PCR, and only the csuE and blaOXA-51-like sequences were amplified in the IC I M-PCR, which could be referred to IC III strains. Standard A. baumannii IC type I, II, and III strains were being used as controls. In this experiment, strains with a double negation of IC type I, II, or III were reported as a variant (V) clonal type and labeled as IC0.
Multilocus sequence typing (MLST)
The internal portions of seven housekeeping genes were scanned via MLST: cpn60 (60-kDa chaperonin), fusA (elongation factor EF-G), gltA (citrate synthase), pyrG (CTP synthase), recA (homologous recombination factor), rplB (50S ribosomal protein L2), and rpoB (ribosomal (RNA polymerase subunit B). MLST website (http://www.pasteur.fr/ recherche/ genopole/ PF8/ mlst/ references-Abaumannii.html). The housekeeping genes for the MLST scheme were selected based on their sequence availability in GenBank and in prior studies of the phylogenetic relationships for the genus Acinetobacter and their presence in other MLST schemes available for other bacterial species. PCR primers for amplifying the seven specified genes were chosen from prior research or particularly suited for them. All PCR amplifications were conducted with Promega's Go Taq Green Master Mix under the following conditions: 35 cycles (denaturation at 94.8C for 30 seconds, annealing at 50.8C for 30 seconds, and extension at 72 8C for 30 seconds) were followed by a 2-minute denaturation at 94 8C and a 5-minute extension at 72 8C. According to its protocol, PCR products were immediately purified from the reaction mixture through using QIAquick PCR Purification Kit (Qiagen GmbH, Hilden, Germany). Internal DNA segments of the chosen housekeeping genes, ranging in size like 297 to 633 bp, were sequenced using an ABI Prism 377 sequencer and the ABI Prism BigDye Terminator Cycle Sequencing Ready Reaction Kit v.3.1. (PE Applied Biosystems, Foster City, CA) in line with the manufacturer's instructions on both strands, PCR primers were utilized for sequencing then ClustalW was used to align the sequence data. (http://www.ebi.ac.uk/Tools/msa/clustalw2/).
Statically analysis
Identification of STs in our isolates due to seven housekeeping genes was selected within the pub MLST. in our search was selected 8 isolates from each ICs (IC0/A11, ICI/A6,48, ICII/A33,50,19 and ICIII/A1,36) according to global lineage Hierarchical dendrogram (Fig. 3) and their resistance patterns (most carbapenem-resistant CR) then were submitted in the MLST Pasteur scheme as in the table (1)
RESULTS
Anti-biotypes analysis of the collection samples
Fifty isolates were detected as A. baumannii, then were differentiated, selective and modified media cultures accompanied by biochemical tests then confirmed by genotypic detection were collected from different hospitals samples sources.
The stander disc diffusion method (antibiotype) was conducted to determine the various resistance patterns versus all antibiotic groups. Classify all isolates according to two bases the first antibiotype to four groups dependent on resistance to carbapenem classes (IMI, MEM & DOR) (C0 30%, C1 6%, C2 34% & C3 30%) and second on three groups dependent on resistance to all antibiotic groups (AB R) or some of them to MDR 26% (13 isolates), XDR 66% (33 isolates) and PDR 8% (4 isolates).
Species diversity and antimicrobial susceptibility
The following steps used sensitivity test for all classes of antibiotics as for penicillin group 80% (AMX&AMC respectively), 74% (PRL), 70% (TIM, SAM & TICER respectively) secondly for cephalosporin group 98% (CRO), 88% (CTX), 74% (CAZ) thirdly for carbapenem 70% MEM, 64% (DOR, 32% (IMI) then for aminoglycoside group 76% (GEN), 68% (AK & TOB respectively), for fluoroquinolone 74% (CIP), 62% (LEV), 50% (GATI), followed for glycylcycline 76% (TE), 50% (TG), 38% (DXT), and 74% (37/50) for polymyxin B, finally the sulfa drug represent 4% sensitive and 96% resistance rate. But still, all isolates gave sensitive results against colistin, except PDR isolates (A11, A22, A25 and A33). Tri-locus multiplex PCR was applied to all isolates to determine which clones belonged via two primers groups with their designated conditions 15, 16, 17. For this purpose were targeted three intrinsic genes, "ompA, csuE & blaOXA-51-like" show in Fig. (1), in our strains' work, were clustered into four groups G1, G2, G3 and G0 where the all strains belonging to the G1 clonal complex yielded expressing ompA csuE and blaOXA-51-like alleles, so provided all three fragments in the PCR, but none in the G2 PCR, whereas strains belonging to G2 showed the predicted opposite findings but in case of G3 clone harboring csuE and blaOXA-51-like alleles from G1with sharing ompA allele as G2 or only have ompA allele as G2 as the report was done by Turton et al.17, But the finding of some outbreak strains not had any set of these genes despite positive to blaOXA-51-like, therefore, be readily assigned as G0 in which that not recording until now by any research. Thirty-seven (74%) different amplification patterns have been allocated to ICs, the widest dissemination of which is G1 (19/50, 38%), followed by G2 (10/50, 20%). The lower dissemination one is ICIII when reached to 16% (8/50), while IC0 reached 26% (13/50) in our study also focus on carbapenem group and other types of antibiotics resistance of isolates (ICs) with their dissemination which were summarized in Fig. (2).
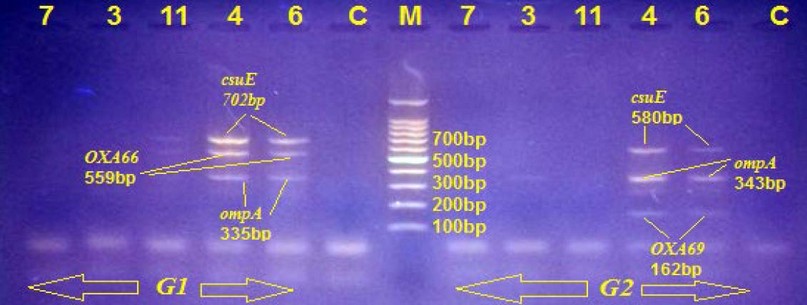
Figure 1. Genetic detection of global lineage by multiplex PCR for two listed primers G1 ompA (335bp), csuE (702bp) & blaOXA-51-like/ 66 (559bp) and G2 ompA (343bp), csuE (580bp) & blaOXA-51-like/66 (162bp). Agarose Gel Electrophoresis (1% Agarose, 7 V/Cm2) and Ethidium Bromide Staining to complete detection. In the middle lane, molecular size DNA Ladder (100bp DNA Ladder) and C refer to the negative control, isolates 4 & 6 under G1 (ICI) because harboring all G1 genes, but 11 & 7 absent any types of G1 or G2, so labeled as G0 (IC0) then isolate no. 3 under G3 (ICIII), which is positive to ompA in G2, absent all G1 genes.
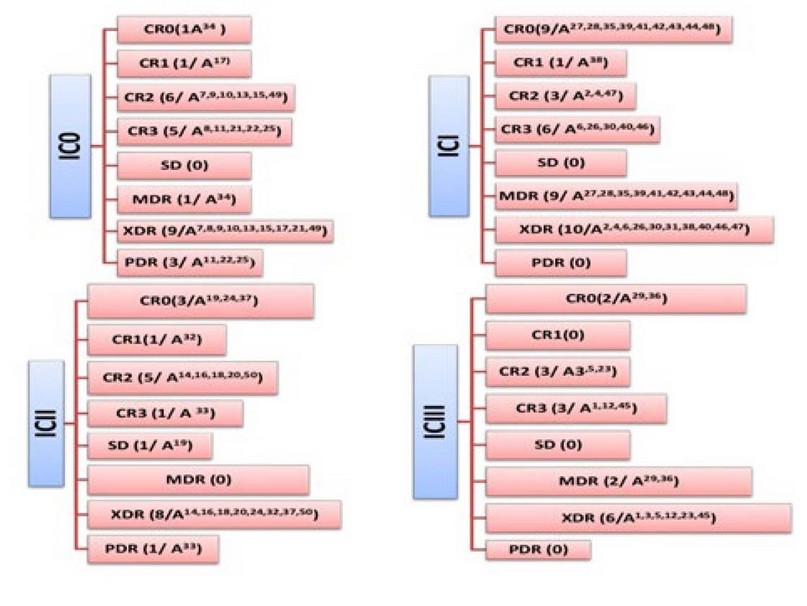
Figure 2. Dissemination of resistances patterns (AR & CR) and ICs.
Phylogenetic analysis
According to the global lineage Hierarchical dendrogram (fig. 3) and their resistance patterns (most carbapenem-resistant CR) our isolates were selected and submitted in MLST Pasteur scheme as in table (1). The relatedness between two strains in Iraq or with the entire world can then be inferred mediated by the allelic profiles varies and illustrated in the following phylogenetic trees.
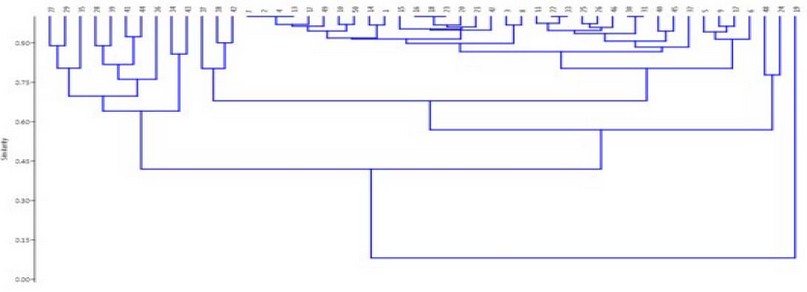
Figure 3. global lineage Hierarchical dendrogram of all currently isolates with their seven clusters and unique A19 isolates as origin strain.

Table 1. submit our isolates in MLST scheme.
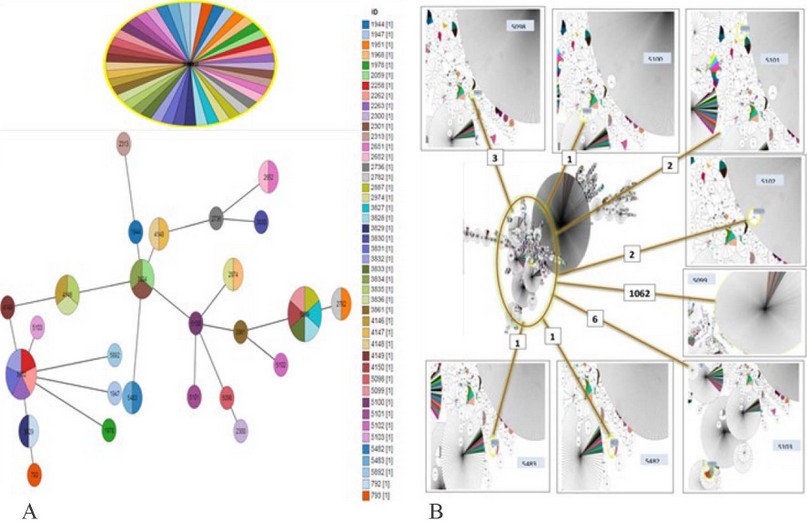
Figure 4. phylogenic tree. A/ MLST diagram shows the relatedness of all isolates in this investigation. STs are shown as filled circles, with the size indicating representation within the dataset. Large circles indicate the primary founder of the MLST clonal complex, small circles indicate sub-group founder within Iraqi only. B/ distribution entire the world.
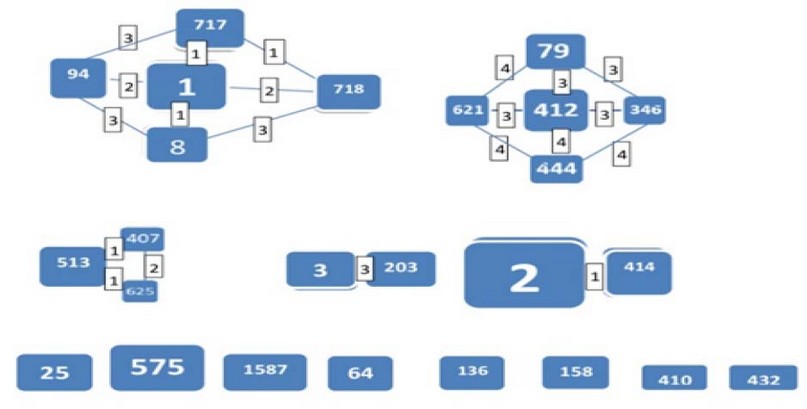
Figure 5. eBURST analysis to clonal complex (CCs). Each square signifies the sequence type and the larger size of each square corresponding to higher frequency of occurrence. Each line indicates that the connected squares share the similar alleles (SLV, DLV & TLV). In the below drawing show related less singleton squares.
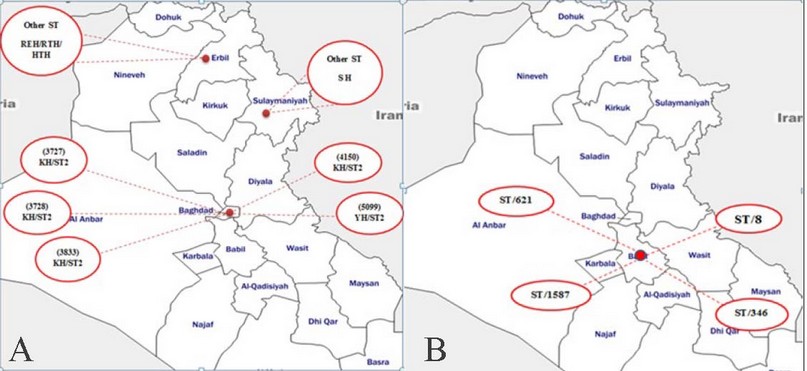
Figure 6. A, recorded returned in Iraq and B, newer recorded in Iraq only.
KH: khadhimain Hospital, YH: Yarmouk Hospital, SH: Sulemani Hospital.

Figure 7. Geographical dissemination of recorded ST entire the world. When ST2 recorded in 1062 isolates from 34 different countries, followed by ST8 were previously reported in six isolates consequently ST 444 (3), ST621 (3), ST 346 (2) and ST 1587 (3).
MLST diagram was done to illustrate the relatedness of all isolates in this investigation locally within Iraq or entire the world as shown in Fig. (4 A& B) than were all particular strains analysis by eBURST to the clonal complex which recorded previously in Iraqi hospitals to determine the epidemiology of STs within this area and within all countries also were determined the originality of Iraqi strains as in Fig. 5, 6 (A & B) and 7 respectively.
DISCUSSION
Reveal the fundamental process that underpins A. baumannii infections, including the evolution of infecting strains, resistance patterns, and genes crucial for resistance development, is essential for the development of viable infection control measures and more efficient medical interventions. Firstly, the isolates were genetically established to reduce the time and effort, according to genus and species-specific genes (16srRNA & oxa 51 respectively). All the isolates must give positive results, indicating all were A. baumannii. These findings support those of other studies and are confirmed by additional methods similar to the current study 18. The most effective antibiotic is colistin, since the percentage of resistance is 8%, four isolates within the PDR group only present this resistance (A11, A22, A25 and A33). Maspi et al., 19 reported 71% of XDR A. baumannii isolates from Iran which convergent with our research, while Kalal et al., 20 reached 80% isolates as XDR rather than 66% as such in the current study. And the last one identified 15/15 MDR isolates with a percentage exceeding 100%, wherein this study detected 13/50 isolates as MDR with a 26% percentage only. As mentioned before Guo et al., 21, 16/67 (23.88%) and 46/67 (68.66%) isolates were defined as MDR and XDR; respectively, these results are acceptable with the current study. The current study results are comparable with the results of Shafigh et al. 22, who identified that more affected on their isolates are colistin but, the percentage of resistance overshoot 30% while in our study does not exceed 8%. But, AL-Kadmy et al. recorded one isolate resist to polymyxin B. These results are more compatible with the search done by Anne et al., 23 for the COL (0%), MEM (83%), CRO (100%), AK (50%), GEN (84%), TE (64%), TG (48%) and SXT (100%) while incompatibles for the IMI (81%), CTX (100%), CAZ (100%), PRL (100%), TOB (87%), CIP (89%) and LEV (91%). So study revealed that the majority of isolates of A. baumannii tested were progressively resistant to approximately all classes of antibiotics may probably be due to the heavy selection pressure from overuse of the antibiotics within the hospital's environment, causing more spreads of these isolates to become breakouts entire the world. Although, increase this resistance may be a takeoff from the recombination by directly acquired from mobile genetic elements (Tn2006 and ISAba1, 2, 3, 4) or harboring to carbapenemase genes (OXA 23, 51, 58 etc.) which can hydrolyze carbapenems and confer resistance to virtually all ß-lactam antibiotics or another mode of resistance patterns as efflux pump correlated with other secretion systems (may be QS or CDI) to confirm their persistence with verity conditions 21, 23, 24.
To better understand bacteria's epidemiology and population structure, the genotypic diversity of strains has really been uncovered using just a wide range of approaches 25, 26. To recognize a limited origin number of clones which responsible for propagation in many countries the Strains of A. baumannii from geographically and genetically diverse European hospitals had been used to distinguish three primary groupings of epidemic strains, which were dubbed European international clones I, II, and clone III with their subtypes from other genotypes remains to be established among themselves 3, 27. Especially during recent years, hospitals in Iraq have experienced numerous outbreaks caused by widespread genotypes of nominee bacteria 28. Enable infection control professionals by this information is also essential for developing and balancing control strategies to reduce the spread and disease burden of A. baumannii in the hospital setting as in Iraq 29, 30. The patterns of groups 0, 1, 2, and 3 identified using this method were found to pertain to IC I, IC II, and IC III, respectively, except IC0 31 as in the report of Hamidian & Nigro 32 were recording more distribution two major clones I & II on a large scale whilst these results were spaced out from results in Thailand by Khuntayaporn et al. 30 where recording ICII high prevalent rather than ICI; the reason may be due to geographical disparity or others.
In previous statistical analysis (Fig. 3) of correlation between these isolates appeared as a cluster with other PDR isolates (22 & 33); give this isolates some solitary, below Hierarchical clustering dendrogram where represented seven clusters (C) in which that A19 was considered the phylogenetic origin of all isolates in this evolutionary tree maybe this was the reason for its being SD then will be divided to two major clusters MDR and XDR/PDR with some exceptions consequently multiple clusters with different characters where C1, C2, C3 and C4 (A27&A29), (A28&A39), (A41&A44) and (A34&A43) respectively came from the one same original branch where all our isolates most MDR. C5 (A38 & A42) when two of them were ICI and standing to same antibiotic except two only (MEM and DXT), otherwise C6 (all other isolates except C7) in this cluster the main and only characteristic of all isolates were its considered as XDR but, exception four PDR isolates within it because its sharing other properties with other isolates in this cluster especial for A22 & A33 as sisters, lastly for C7 (A5, A6, A9 and A12) which all of it were XDR and have resistance to 21from 24 types of antibiotics.
We also focus on carbapenem resistance isolates, where only two from fifty isolates ICI and ICII (A38 & A32) have a MEM resistance only. However, most of ICI unable to stand against this group despite this most of them recorded MDR and XDR rather than other types on the contrary IC0 most of them standing except A34 and most of them considered as PDR these results it's opposite to Zhang et al., 33 reports, but ICII and ICIII have variable results in these cases. ICII have only one PDR isolate A33 and one SD isolate A19; in the thesis of Dahdouh 34 illustrated 37/42 under ICII and have carbapenem resistance while in our study resistance ICII isolates detected in 7/10; consequently, either PCR assays or searches of genomic data targeting the most discriminatory Global clone could simplify the analysis of the epidemiological studies of outbreaks involving multiple closely related types.
The performance of molecular typing in this study was supported by MLST analysis; this pattern has been adopted in approximately 10 researches in Iraq, as in Fig. 4 (A) in which Iraqi isolated from a variety of sources (16 wounds, 12 blood, 6 skin, 5 sputa, 2 CSF, 2 environments, 1 urine and 1 other) with the various types of MLST (Oxford/Pasteur 24 and only Pasteur 21) (http://pubmlst.org/abaumannii/). At the same time, the study comprised geographical dissemination of these isolates in different geographic regions worldwide (recorded 6331 isolates) as in Fig. 4 (B). Based on the phylogenetic data, ST2 recorded in 1062 isolates from 34 different countries (5 Unknown, 1 Germany, 1 South Africa, 1 Ireland, 1Portugal, 1 Israel, 2 Poland, 2 Australia, 2 Lebanon, 2 Egypt, 2 Belarus, 3Lithuania, 11thailand, 11France, 11 Italy, 3 Taiwan, 7 Greece, 8 Japan, 5 Malaysia, 5 South Korea, 6 Iraq, 18 Singapore, 7UK, 7 Sweden, 5 Czech Republic, 8 The Netherlands, 6 Spain, 6 Norway, 3 Denmark, 19 Croatia, 26 Saudi Arabia, 32 Turkey, 91 Russia, 348 China, 397 USA) in which that indicator to variability genome of strain type, on the contrary, the rest had very few repeaters as six Europeans for ST8 were previously reported 1, 1, 3 and 1 in the Netherland (lower respiratory tract LRT), Italy (blood), Greece (2 LRT & 1 urine) and Turkey (blood) respectively.
For the more comprehensive analysis of the possible patterns of evolutionary descent, a set of rules were proposed and implemented in the eBURST algorithm, which allows the division of a data set into several clusters of related strains, dubbed clonal complexes (CC), by implementing a simple model of clonal expansion and diversification 35, 36. Within each clonal complex, the rules identify which links between STs, in our study founding that isolates could cluster to more than five with Iraqi isolates as in the previously Fig. 5 and the major of isolates exhibited closely relationships in the evolution with most major clonal complex belong to CC2 were highly frequent in Iraq as MLST scheme dataset as found to be a dominant clone in all this region while our study founding only one from 8 isolates (A6) under ST2 and ICI. But ST1 was not among our isolates also within CC3& CC5. In 2005, only a single ST2 isolate was obtained in Iraq then increased progressively until 2016 when recorded eight isolates with six ST were recorded 3 & 2 for ST2 &ST1 respectively, and only one new ST was recorded (ST203) by Qasim et al. 37 for our time this indicating local distribution over time. Noted, the molecular characterization of the isolates by MLST identified 4 new different STs (8, 444, 346 & 621) within Iraq, indicating a high genotypic diversity and possible divergent evolution of these strains from their original clones 37, 38 in which firstly within CC1, it has four ST (1, 8*, 94, 717 and 718) and ST8 represent within IC0 follow in the CC2 (ST2* & ST414); containing as previous explain the predicted ancestor ST2 under ICI and high prevalence one in Iraq which also approved by Qasim et al 37 reports. CC4 involve ST79, ST346*, ST412, ST444* and ST621* so in this complex, a three newer recorded types in this country with lower connected to other STs in their CC and considered as sub-group where its A33ST346, A36ST621 and A48ST444 isolated from Medical Center Hospital for first two types and Al-Yarmouk Hospital/Baghdad for the last one which illustrated in Fig. (6). While there, only three of our isolates were singleton (A1ST136*, A11ST1587 & A191587). HA-1 (ST 136/ICIII) was recorded previously in Kurdistan/Iraq were isolated from different patients; most of them burns or ICU cases in their hospitals (in Erbil & Sulaimania) and clear up the successful clones to spreading among different cities and countries 39.
Our strain A50 ICII was isolated from CSF source in which lower cases in the world were isolated from this source because it a more difficult to obtain its. And two strains were isolated in 2012 from urine of patients in Japan hospital which have closely relatedness to A48 (ST444/ICI) otherwise A36 (ST621/ICIII) were obtained from blood and a grouped together with France clone (unknown sources), in spite of that our study locate three clones (A33,19,11/ ST346 & ST1587 respectively) have only one along the universal clones firstly one isolated from Switzerland food at 2013 and the two last clone from Germany environment after our record (WWTP-AS-0520-A-V-5) that indicated some Iraqi clones originated from nonclinical samples as notably in the A. baumannii ST2 has been extensively isolated from humans, while some of the recent reports have also indicated the transmission of ST2 by domestic animals to human which may be served as reservoirs as in A33 (id: 5100) which founding that only two isolates in the world as ST346 and in Switzerland the source of second isolate (id: 1856) was chicken, particularly CR, due to their selective advantage compared to the susceptible strains as in our A19 strain 40, 41. As for the last (A19,11) have a unique clone, especially A19 may be considered as a wild type when acquired their resistance by alteration or modifications due to mutation and recombination in their genome as adaptability within the varies environment to become the phylogenetic origin for the rest of the isolates branching from them.
On the other hand, could be found some types of ST entire the world correlated with antibiotic and resistance genes where ST2 in our study produce full resistance toward three major antibiotic groups (aminoglycoside/ fluoroquinolone/ carbapenem) as recorded ZQ1,2,7 in AL- khadhimai Hospital/ Baghdad/ Iraq (id/ 3827,3828 & 3833) when resistance to the same types of antibiotics give suggesting that it's maybe one of them was originally from the other or acquired its character from same exposure geographic areal conditions within the Iraqi hospitals to give this similarity. For newly ST recorder, the emergence of such strains in Iraq may predict an increase in the risk of how to deal with it and reduce it where it could be seen that in two PDR strains A33 ST 346 & A11 ST 1587 under ICII and IC0, respectively with resistance to all types of carbapenem group also A1ST, A6 ST 2 & A50 ST 8 represent a more resistance rate to antibiotic groups (XDR) except two only (polymyxin & tetracycline groups) otherwise two newer MDR strains in Iraq A36 & A48 which may be originated from France and Japan respectively. The distribution of the resistance, especially against carbapenem across multiple genetic lineages, was observed both at the national and local levels suggest an expansion of adaptations in Iraq is driven by the horizontal dissemination of resistance determinants across diverse genetic backgrounds rather than by the sole expansion of single resistant genetic clones, and that resistance to multiple antibiotics is acquired en bloc via MGEs 42. At the local and global scale, our study identifies several high-risk clones of A. baumannii ST 346 & ST 1587 in Iraq by linking clonal relatedness, geographic clustering, epidemiological data with the possibility of frequent international transmission. Finally, newer recorded ST in strain A1 around the world indicated more modification and alteration in their genomic sequences of Iraqi isolates to become origin to other strains in other different countries.
CONCLUSION
In conclusion, it shows high rates of CRAB isolates and predominance of XDR and PDR under ICs in Iraqi strains compared with previous years. These findings suggest a strong correlation between resistance genes and ICs types determined on a local since different associations were detected in the different Baghdad hospitals. Also, epidemiological of Iraqi clones worldwide with newly recorded types may be more risk factors for epidemic distribution of the chronic diseases associated with these bacteria.
Acknowledgment
The authors would like to express their gratitude to Mustansiriyah University (http://uomustansiriyah.edu.iq/) in Baghdad, Iraq, for their assistance with this project.
REFERENCE
- Castillo-Ramírez, S., & Graña-Miraglia, L. Inaccurate multilocus sequence typing of Acinetobacter baumannii. Emerging infectious diseases, 2019. 25(1), 186.
- Hu, Y. F., Hou, C. J. Y., Kuo, C. F., Wang, N. Y., Wu, A. Y. J., Leung, C. H., ... & Yeh, H. I. Emergence of carbapenem-resistant Acinetobacter baumannii ST787 in clinical isolates from blood in a tertiary teaching hospital in Northern Taiwan. Journal of microbiology, immunology and infection, 2017. 50(5), 640-645.
- Gaiarsa, S., Batisti Biffignandi, G., Esposito, E. P., Castelli, M., Jolley, K. A., Brisse, S. et al. Comparative analysis of the two Acinetobacter baumannii multilocus sequence typing (MLST) schemes. Frontiers in microbiology, 2019. 10, 930.
- Zhang, Y. Y., Liang, Z. X., Li, C. S., Chang, Y., Ma, X. Q., Yu, L., & Chen, L. A. Whole-Genome analysis of an extensively drug-resistant Acinetobacter baumannii strain XDR-BJ83: insights into the mechanisms of resistance of an ST368 strain from a Tertiary Care Hospital in China. Microbial Drug Resistance, 2018. 24(9), 1259-1270.
- Redfearn, D. P., Trim, G. M., Skanes, A. C., Petrellis, B., Krahn, A. D., Yee, R., & Klein, G. J. Esophageal temperature monitoring during radiofrequency ablation of atrial fibrillation. Journal of cardiovascular electrophysiology, 2005. 16(6), 589-593.
- Giannouli M, Cuccurullo S, Crivaro V, Di Popolo A, Bernardo M, Tomasone et al. Molecular epidemiology of multidrug-resistant Acinetobacter baumannii in a tertiary care hospital in Naples, Italy, shows the emergence of a novel epidemic clone. J. Clin. Microbiol. 2010. 48:1223–1230. http://dx.doi.org/10 .1128/JCM.02263-09.
- Zarrilli R, Pournaras S, Giannouli M, Tsakris A. Global evolution of multidrug-resistant Acinetobacter baumannii clonal lineages. Int. J. Antimicrob.2013.Agents,41:11–19. http://dx.doi.org/10.1016/j.ijantimicag.2012 .09.008.
- Hamidian, M., Hawkey, J., Wick, R., Holt, K. E., & Hall, R. M. Evolution of a clade of Acinetobacter baumannii global clone 1, lineage 1 via acquisition of carbapenem-and aminoglycoside-resistance genes and dispersion of ISAba1. Microbial genomics, 2019. 5(1).
- Schultz, M. B., Thanh, D. P., Do Hoan, N. T., Wick, R. R., Ingle, D. J., Hawkey, J. et al. Repeated local emergence of carbapenem-resistant Acinetobacter baumannii in a single hospital ward. Microbial genomics, 2016. 2(3).
- Holt, K. E., Kenyon, J. J., Hamidian, M., Schultz, M. B., Pickard, D. J., Dougan, G., & Hall, R. M. Corrigendum: Five decades of genome evolution in the globally distributed, extensively antibiotic-resistant Acinetobacter baumannii global clone 1. Microbial genomics, 2019. 5(7).
- Correa, A., Del Campo, R., Escandón-Vargas, K., Perenguez, M., Rodríguez-Baños, M., et al. Distinct genetic diversity of carbapenem-resistant Acinetobacter baumannii from Colombian hospitals. Microbial Drug Resistance, 2018. 24(1), 48-54.
- Evans, B. A., & Amyes, S. G. OXA β-lactamases. Clinical microbiology reviews, 2014. 27(2), 241-263.
- Pearce, M. E., Chattaway, M. A., Grant, K., & Maiden, M. C. A proposed core genome scheme for analyses of the Salmonella genus. Genomics, 2020. 112(1), 371-378.
- CLSI. M100-S25 Performance Standards for Antimicrobial Susceptibility Testing. Thirty one Informational Supplement. Clinical and Laboratory Standards Institute, 2021.
- Woodford, N., Ellington, M. J., Coelho, J. M., Turton, J. F., Ward, M. E., Brown, S. et al. Multiplex PCR for genes encoding prevalent OXA carbapenemases in Acinetobacter spp. International journal of antimicrobial agents, 2006. 27(4), 351-353.
- Turton, J. F., Ward, M. E., Woodford, N., Kaufmann, M. E., Pike, R., Livermore, D. M., & Pitt, T. L. The role of IS Aba1 in expression of OXA carbapenemase genes in Acinetobacter baumannii. FEMS microbiology letters, 2006. 258(1), 72-77.
- Turton, J. F., Gabriel, S. N., Valderrey, C., Kaufmann, M. E., & Pitt, T. L. Use of sequence‐based typing and multiplex PCR to identify clonal lineages of outbreak strains of Acinetobacter baumannii. Clinical microbiology and infection, 2007. 13(8), 807-815.
- Al-Kadmy, I. M. S., Ali, A. N. M., Salman, I. M. A., & Khazaal, S. S. Molecular characterization of Acinetobacter baumannii isolated from Iraqi hospital environment. New microbes and new infections, 2018. 21, 51-57.
- Maspi, H., Hosseini, H. M., Amin, M., & Fooladi, A. A. I. High prevalence of extensively drug-resistant and metallo beta-lactamase-producing clinical Acinetobacter baumannii in Iran. Microbial pathogenesis, 2016. 98, 155-159.
- Kalal, B. S., Chandran, S. P., Yoganand, R., & Nagaraj, S. Molecular characterization of carbapenem-resistant Acinetobacter baumannii strains from a tertiary care center in South India. Infectio, 2020. 24(1), 27-34.
- Guo, T., Shan, C., Li, J., Feng, H., Li, M., Zheng, W., ... & Bao, G. Correlation between antibiotic resistance and serum resistance in Acinetobacter baumannii. INTERNATIONAL JOURNAL OF CLINICAL AND EXPERIMENTAL MEDICINE, 2019. 12(8), 9804-9814.
- Shafigh, M., Rajabnia, R., Yahyapour, Y., Shahandashti, E. F., Khafri, S., & Namvar, A. E. Evaluation of Aminoglycoside Resistance Genes in Acinetobacter Baumannii Isolated from Different Parts of Babol Hospitals. Biomed J Sci & Tech Res. 2018. DOI: 10.26717/BJSTR.2018.08.001675. ISSN: 2574-1241.
- Anane, Y. A., Apalata, T., Vasaikar, S., Okuthe, G. E., & Songca, S. Molecular Detection of Carbapenemase-Encoding Genes in Multidrug-Resistant Acinetobacter baumannii Clinical Isolates in South Africa. International Journal of Microbiology, 2020.
- Botelho, J., Roberts, A. P., León-Sampedro, R., Grosso, F., & Peixe, L.. Carbapenemases on the move: it’s good to be on ICEs. Mobile DNA, 2018. 9(1), 1-11.
- Dijkshoorn, L., Aucken, H., Gerner-Smidt, P., Janssen, P., Kaufmann, M. E., Garaizar, J., ... & Pitt, T. L. Comparison of outbreak and nonoutbreak Acinetobacter baumannii strains by genotypic and phenotypic methods. Journal of clinical microbiology, 1996. 34(6), 1519-1525.
- Maciel, W. G., da Silva, K. E., Croda, J., Cayo, R., Ramos, A. C., de Sales, R. O., ... & Simionatto, S. Clonal spread of carbapenem-resistant Acinetobacter baumannii in a neonatal intensive care unit. Journal of Hospital Infection, 2018. 98(3), 300-304.
- Van Dessel, H., Dijkshoorn, L., van der Reijden, T., Bakker, N., Paauw, A., van den Broek, P., ... & Brisse, S. Identification of a new geographically widespread multiresistant Acinetobacter baumannii clone from European hospitals. Research in microbiology, 2004. 155(2), 105-112.
- Higgins, P. G., Hagen, R. M., Kreikemeyer, B., Warnke, P., Podbielski, A., Frickmann, H., & Loderstädt, U. Molecular Epidemiology of Carbapenem-Resistant Acinetobacter baumannii Isolates from Northern Africa and the Middle East. Antibiotics, 2021. 10(3), 291.
- Tomaschek, F., Higgins, P. G., Stefanik, D., Wisplinghoff, H., and Seifert H. Head-to-Head Comparison of Two Multi-Locus Sequence Typing (MLST) Schemes for Characterization of Acinetobacter baumannii Outbreak and Sporadic Isolates. PLoS One, 2016. 11(4), e0153014.
- Khuntayaporn, P., Kanathum, P., Houngsaitong, J., Montakantikul, P., Thirapanmethee, K., & Chomnawang, M. T. Predominance of international clone 2 multidrug-resistant Acinetobacter baumannii clinical isolates in Thailand: a nationwide study. Annals of clinical microbiology and antimicrobials, 2021. 20(1), 1-11.
- Karah, N., Haldorsen, B., Hegstad, K., Simonsen, G. S., Sundsfjord, A., & Samuelsen, Ø. Species identification and molecular characterization of Acinetobacter spp. blood culture isolates from Norway. Journal of antimicrobial chemotherapy, 2011. 66(4), 738-744.
- Hamidian, M., & Nigro, S. J. Emergence, molecular mechanisms and global spread of carbapenem-resistant Acinetobacter baumannii. Microbial genomics, 2019. 5(10).
- Zhang, X., Li, F., Awan, F., Jiang, H., Zeng, Z., & Lv, W. Molecular epidemiology and clone transmission of carbapenem-resistant Acinetobacter baumannii in ICU rooms. Frontiers in cellular and infection microbiology, 2021. 11.
- Dahdouh, E. Epidemiological and molecular analysis of virulence and antibiotic resistance in Acinetobacter baumannii. 2017 (Doctoral dissertation, Universidad Complutense de Madrid).
- Feil, E. J., Li, B. C., Aanensen, D. M., Hanage, W. P., & Spratt, B. G. eBURST: inferring patterns of evolutionary descent among clusters of related bacterial genotypes from multilocus sequence typing data. Journal of bacteriology, 2004. 186(5), 1518-1530.
- Francisco, A. P., Bugalho, M., Ramirez, M., & Carriço, J. A. Global optimal eBURST analysis of multilocus typing data using a graphic matroid approach. BMC bioinformatics, 2009. 10(1), 1-15.
- Qasim, Z. J., Kadhim, H. S., & Abdulamir, A. S. Identification of Antibiotic Resistance Genes in Multi-Drug Resistant Acinetobacter Baumannii Clinical Isolates of Iraqi Patients (Zq Strains), Using Whole-Genome Sequencing. INTERNATIONAL JOURNAL OF PHARMACEUTICAL AND CLINICAL RESEARCH, 2019. 10(04), 670-680.
- Ou, H. Y., Kuang, S. N., He, X., Molgora, B. M., Ewing, P. J., Deng, Z., ... & Xu, H. H. Complete genome sequence of hypervirulent and outbreak-associated Acinetobacter baumannii strain LAC-4: epidemiology, resistance genetic determinants and potential virulence factors. Scientific reports, 2015. 5(1), 1-13.
- Ganjo, A. R., Maghdid, D. M., Mansoor, I. Y., Kok, D. J., Severin, J. A., Verbrugh, H. A., ... & Goessens, W. OXA-carbapenemases present in clinical Acinetobacter baumannii-calcoaceticus complex isolates from patients in Kurdistan region, Iraq. Microbial drug resistance, 2016. 22(8), 627-637.
- Püntener-Simmen, S., Zurfluh, K., Schmitt, S., Stephan, R., & Nüesch-Inderbinen, M. Phenotypic and genotypic characterization of clinical isolates belonging to the Acinetobacter calcoaceticus-Acinetobacter baumannii (ACB) complex isolated from animals treated at a veterinary hospital in Switzerland. Frontiers in veterinary science, 2019. 6, 17.
- Taj, Z., Rasool, M. H., Almatroudi, A., Saqalein, M., & Khurshid, M. Extensively drug-resistant Acinetobacter baumannii belonging to international clone II from a pet cat with urinary tract infection; the first report from Pakistan. Polish journal of microbiology, 2020. 69(2), 231.
- Argimón, S., Masim, M. A., Gayeta, J. M., Lagrada, M. L., Macaranas, P. K., Cohen, V., ... & Carlos, C. C. Integrating whole-genome sequencing within the National Antimicrobial Resistance Surveillance Program in the Philippines. Nature communications, 2020. 11(1), 1-15.
Received: 1 December 2021 / Accepted: 15 January 2022 / Published: 15 February 202
Citation: Kadhom HA, Ali MR. Epidemiological Molecular Analysis of Acinetobacter baumannii isolates using a multilocus sequencing typing and Global lineage. Revis Bionatura 2022;7(1). 29. http://dx.doi.org/10.21931/RB/2022.07.01.29
