2022.07.02.40
Files > Volume 7 > Vol 7 No 2 2022
1 Dept. of Med. and Mol. Biotech., College of Biotechnology, Al-Nahrain University, Baghdad, Iraq; [email protected];
2 Dept. of Med. and Mol. Biotech., College of Biotechnology, Al-Nahrain University, Baghdad, Iraq; [email protected];
3 DNA forensic Center for Research and Training, Al-Nahrain University, Baghdad, Iraq 2; [email protected]
* Correspondence: e-mail; [email protected]
Available from: http://dx.doi.org/10.21931/RB/2022.07.02.40
ABSTRACT
Multidrug-resistant (MDR) and pathogenic E. faecium is a crisis in healthcare settings. This survey aimed at antibiotic susceptibility profiling and virulence determinants of E. faecium. This study pooled 100 fecal E. faecium isolates identified by phenotypic and molecular tests. Antibiotic susceptibility and ampicillin MIC were determined according to clinical and laboratory standards institute (CLSI) 2017. Moreover, biofilm formation was assayed by a microtiter tissue plate assay. Virulence genes pattern was performed by polymerase chain reaction (PCR) method. Among 460 fecal samples, 100 isolates of E. faecium were identified, among which the highest resistance was related to penicillin (81%), cephalothin (78%), cefazolin (76%), tetracycline and cefepime (69%). In contrast, 83% of them were susceptible to vancomycin. Moreover, four vancomycin-resistant isolates had vancomycin MIC>32µg/mL, and 11 isolates had MIC>8µg/mL. Of 32 ampicillin-resistant isolates, 18 (56%) produced strong biofilm, and 14 isolates (44%) produced moderate biofilm. Among 17 vancomycin-resistant E. faecium (VREfa), 15 (88.23%) isolates produced strong biofilms, and two of them produced moderate-level biofilms, which was significantly higher than susceptible isolates (p=0.0144). The vanA (vancomycin MIC: 16-64µg/mL) and vanB (vancomycin MIC: 8-64µg/mL) genes were detected in twelve and five isolates, respectively. The rate of adhesin genes ace, esp and ebp included 68%, 97% and 82%, respectively. All the VREfa harbored the ace, esp and ebp genes. The antibiotic resistance rate of E. faecium was low. Biofilm formation was significantly different between VREfa and vancomycin-susceptible isolates but not between k9ampicillin-resistant and ampicillin-susceptible isolates. All the VREfa harbored the ace, esp and ebp genes. The virulence adhesin genes were not significantly associated with biofilm formation. Further studies are essential to appreciate better the relation between biofilm formation, drug non-susceptibility and adhesin genes.
Keywords: E. faecium, antibiotic resistance, biofilm formation, virulence genes
INTRODUCTION
In recent years, Enterococcus species have been identified as the major contributing pathogens in hospital infections and community-acquired infections 1,2, due to the acquisition of multiple antibiotic resistance genes, such as beta-lactams, cephalosporins, trimethoprim-sulfamethoxazole and glycopeptides. Some Enterococci are inherently resistant to beta-lactam, and aminoglycoside antibiotics are considered a medical crisis 3-6. Glycopeptides like vancomycin are used as the last line of antibiotics in treating multidrug-resistant (MDR) Gram-positive bacteria 7. Vancomycin-resistant Enterococci (VRE) carrying the resistance genes comprise the most common pathogens in hospitals worldwide 8,9.
In numerous studies, the adhesin and effective binding of Enterococci is reported on the levels of medical devices, such as intravenous and urinary catheters and ocular lenses, and lead to biofilm production 10,11. The Enterococci pathogenic spp are less dependent on their virulence factors and the ability to adapt to environmental conditions 8,10. Their resistance to antibiotics and ecological stresses, binding properties, and biofilm formation has resulted in crucial pathogenic features associated with implant infections 5,6,8. In addition, Enterococci are significant contributors to nosocomial infections, and biofilm formation, which employ a wide variety of factors, including genetic factors, various environmental conditions, and signals mostly unknown and required for identification and characterization 11,12. Enterococci, development of resistance by multiple mechanisms and a series of virulence factors, are due to physical inhibition and biofilm conditions. A complete understanding of genetic and environmental factors involved in the development of biofilms leads to improved and accurate strategies for controlling biofilm formation. This study aimed to investigate some E. faecium antibiotic resistance, biofilm formation, and virulence factors.
MATERIALS AND METHODS
Bacterial isolates
In this study, 460 fecal samples were collected in Baghdad city during May 2017-October 2019. The E. faecium isolates were identified using conventional biochemical (VITEK 2 system, Biom' erieux, Marcy l'Etoile, France) and molecular tests. They were kept in brain heart infusion broth at -70 ° C. The bacterial isolates were cultured and stoked again for further studies.
Antibiotic susceptibility test
The isolates were tested using Kirby Bauer disk diffusion and Muller Hinton agar medium by suspension equivalent to the half MacFarland for susceptibility to discs including penicillin 10U, ampicillin (10μg), cephalothin (30μg), vancomycin (10µg), tetracycline (30µg), erythromycin (30µg), amikacin (10µg), cefazolin (10μg), ceftriaxone 30μg) and cefepime (30μg) [MAST, UK]. After incubation, the diameter of the inhibition zone was measured according to CLSI 2017 standards 11.
The minimum inhibitory concentration
The minimum inhibitory concentration (MIC) for ampicillin and vancomycin was determined using the agar dilution method onto the medium of brain hearth infusion agar (BHI). In this study, the standard strain of E. faecalis (ATCC 292152) was used as a control, and the results were interpreted following the CLSI 2020 protocol 11.
Biofilm formation
At this stage, the phenotypic measurement of the biofilm formation was carried out using a 96-well microplate 13.
Extraction of total genomic DNA
For DNA extraction, the bacterium was cultured in a BHI medium (Merk, Germany) for 24 hours at 37 ° C in an incubator. Then according to the DNA purification kit purchased from Cinapure DNA, the DNA was extracted from isolates. Quantification of extracted DNA was conducted using a spectrophotometer 14.
Polymerase chain reaction
An investigation of the presence of a specific gene in E. faecium was conducted using specific primers in Table1. The molecular identification of isolates was performed by amplifying the ddl gene with a 658bp size.
Additionally, virulence and vancomycin resistance genes in E. faecium, including esp, ace, ebpA, were investigated using specific primers depicted in table1. Moreover, for vancomycin resistance genes, van A primers included F: ATCAACCATGTTGATGTAGC and R: AAGGGATACCGGACAATTCA. The van B primers included F: ACCCTGTCTTTGTGAAG and R: GAAATCGCTTGCTCAAT 8.
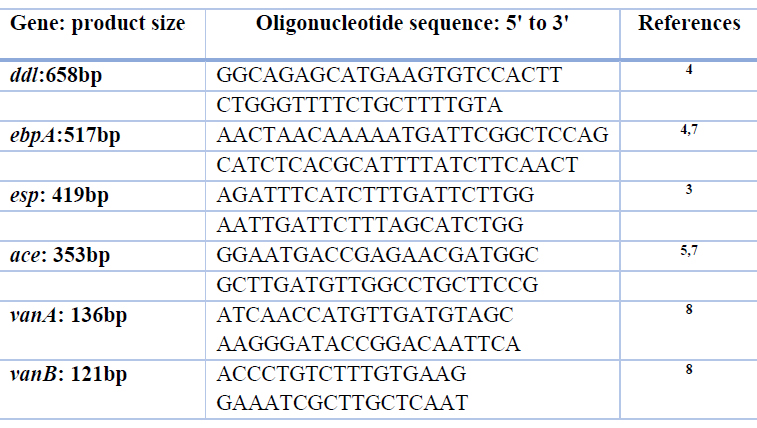
Table1. Primers sequences used in this study
Statistical analysis
Comparisons of the prevalence of antibiotic resistance and virulence genes between groups were performed using the SPSS version 20 (Chicago, IL, USA). The statistical tests included Chi-Square or Fisher's exact test, and ANOVA, where a p value<0.05 was a significant result.
RESULTS
E. faecium antibiotic resistance pattern
Among the isolates of E. faecium, the highest resistance was against the penicillin (81%), cefazolin (76%), tetracycline (69%), and cephalothin (78%), cefepime (69%) and ceftriaxone (67%) and erythromycin (68%). The lowest resistance rate was against ampicillin (32%), vancomycin (21%) and amikacin (21%). Therefore, 64% of isolates were simultaneously resistant to tetracycline, erythromycin and β-lactams and were considered as MDR E. faecium.
Figure 1. The antibiotic susceptibility test pattern
MIC of ampicillin and vancomycin
The MIC of ampicillin among 32% of E. faecium included ≥16 in the resistance range. The MIC in 20% of isolates for this antibiotic was < 8 μg /mL. Moreover, the vancomycin MIC ranged 2-64µg/mL. Indeed, four isolates had MIC>32µg/L, and 11 had MIC>8µg/mL (table2).
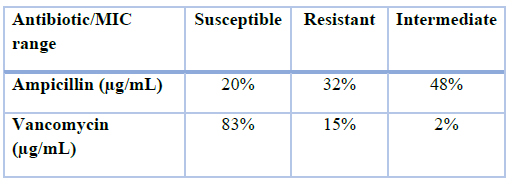
Table2. The MIC levels of vancomycin and ampicillin
Biofilm formation
Phenotypic biofilm formation using a 96-well microplate assay is as followings:
Among 68 isolates of ampicillin-susceptible E. faecium, 30 isolates (44%) formed strong biofilms, and 38 isolates (56%) formed moderate-level biofilms. Furthermore, among 32 ampicillin-resistant isolates, 18 (56%) of them produced moderate-level biofilm and 14 isolates (44%) did not produce biofilm. Among 17 vancomycin-resistant E. faecium (VRE), 15 (88.23%) isolates produced strong biofilms and two of the produced moderate-level biofilms which was significantly higher than susceptible isolates (p=0.0144). There was no significant difference between MDR and non-MDR isolates regarding biofilm formation.
Molecular tests
PCR reaction exhibited that E. faecium isolates amplified the ddl gene (Figure 2) and biofilm-associated genes (Figure 3). The prevalence of ace, esp, and ebp genes was 68%, 97% and 82%, respectively.
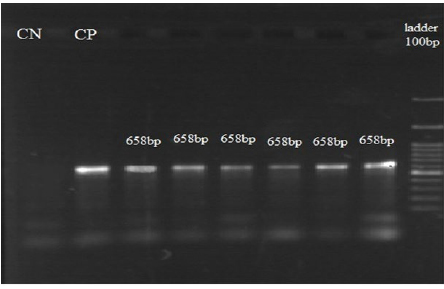
Figure 2. The electrophoresis of ddl gene (658bp), approving samples, CN: negative control, CP: positive control and 100bp DNA ladder have been exhibited.
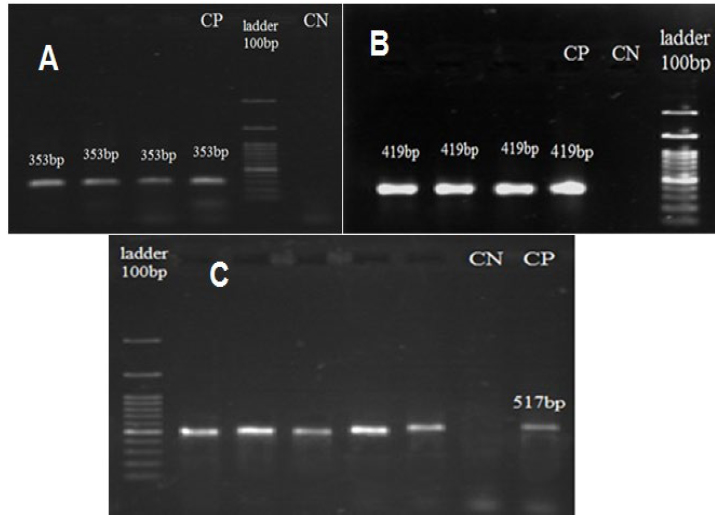
Figure 3. A: The electrophoresis of ace gene (353bp), in which positive samples, CN: negative control, CP: positive control and 100bp DNA ladder have been exhibited. B: The electrophoresis of esp gene (419bp), in which positive samples, CN: negative control, CP: positive control and 100bp DNA ladder have been exhibited. C: The electrophoresis of ebp gene (517bp), in which positive samples, CN: negative control, CP: positive control and 100bp DNA ladder have been exhibited.
Vancomycin resistance genes
The vanA (MIC: 16-64µg/mL) and vanB (MIC: 8-64µg/mL) genes were detected in 12 and five isolates, respectively. Biofilm formation was significantly different between VREfa and vancomycin-susceptible isolates but not between ampicillin-resistant and ampicillin-susceptible isolates. All the VREfa harbored the ace, esp and ebp genes.
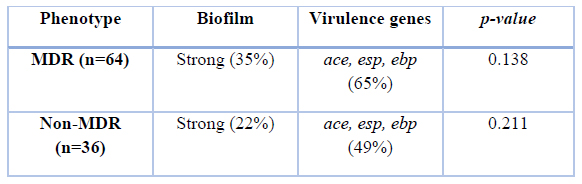
Table 3. The relation between MDR phenotype, biofilm formation and resistance genes
Table 3 exhibited that MDR and non-MDR isolates had no significant difference regarding the virulence rate and biofilm formation using the chi-square test.
DISCUSSION
Although E. faecalis infections are ten times more frequent than those of E. faecium, E. faecium has increased recently, mainly following drug-resistant strains. Enterococci are the second most common opportunistic nosocomial pathogens which can spread antibiotic resistance genes 10,15-17. The ability of Enterococci to colonize the intestinal tract of the individuals admitted for long periods is a critical factor affecting drug resistance. Enterococci act as a source of transmission cycles in the digestive system and release antibiotic resistance 1,2.
The data collected by the ICU surveillance system outlined that E. faecium was the third most common cause of bloodstream infections, the third leading cause of urinary excretion, the most commonly isolated of surgical site infections and the fourth most widely separated of all regions 12,14,18. E. faecium are opportunistic pathogens, and the increasing severity of infections in hospital patients has been associated with E. faecium. Extensive and widespread use of broad-spectrum antibiotics in the hospital facilitates the spread of drug-resistant organisms. The predisposing factors are colonization of Enterococci, hospitalization in the intensive care unit, prolonged use of antibiotics and prolonged hospitalization. Although several antibiotics have been effective against VRE, mortality rates remain high in patients 19. In most cases, due to the excessive use of antibiotics, there are many cases of drug resistance in pathogens, which leads to failure in treatment and the emergence of many complications, despite the high cost of treatment 20,21. Drug resistance to antibiotics in different world regions is additional due to genetic changes in the strains and the difference in the amount of antibiotics 22.
Antibiotic resistance
Herein, the highest resistance rate was against penicillin (81%), cefazolin (76%), tetracycline (69%), cephalothin (78%), and cefepime (69%) and ceftriaxone (67%) and erythromycin (68%). These results indicate that antibiotic resistance to penicillin predisposes the development of resistance to carbapenems and other beta-lactams 23. The lowest resistance rate was against amikacin (32%), vancomycin and ampicillin. Therefore, 64% of isolates were simultaneously resistant to tetracycline, erythromycin and β-lactams and considered as MDR- E. faecium. In a study, the rate of drug resistance was low at 21. Therefore, aminoglycosides are still appropriate for the treatment of enterococcal infections. VREfa rate was 17% and the vanA (MIC: 16-64µg/mL) and vanB (MIC: 8-64µg/mL) genes were detected in 12 and five of them, respectively. Noticeably, the penicillin and ampicillin resistance rate of vanA-type VREfa included 100% for both. Likewise, 96% and 93% of vanB-type VREfa were resistant to penicillin and ampicillin disks, respectively, significantly higher than vancomycin-susceptible isolates.
Biofilm formation
E. faecium is a crucial agent with a vast antibiotic resistance rate and biofilm formation among Enterococci species. The ability to form biofilm on living surfaces is one of the key pathogenicity factors for this bacterium. E. faecium is associated with biofilm formation in endocarditis, urinary tract infections, and dental root and ocular infections. Various factors such as antibiotic resistance and the expression of genes involved in pathogenicity and biofilm formation contribute to the stability of this bacterium in different conditions and the spread of infection 17. According to our study, biofilm formation was significantly different between VREfa and vancomycin-susceptible isolates but not between ampicillin-resistant and ampicillin-susceptible isolates. All the VREfa harbored the ace, esp and ebp genes. Of the studied genes, ebp and esp had the highest rate and ace gene was the lowest among the isolates. 79% of the isolates had the pattern ebp + / esp +. Most antibiotic-resistant isolates had virulence type ebp / esp (78.1%). Gok detected ace (83.6%), esp (66.4%), ebp (60.0%) in Turkey 24, but the rate of these genes has not been reported in this country.
Virulence factors
In numerous studies, the adhesion and effective binding of Enterococci have been reported on the levels of medical devices, such as intravenous and urinary catheters, ocular lenses, and ultimately biofilm production 12,16,17. In a study by Ochoa et al., the esp gene was observed in 83% of isolates, being similar to our result (97%) 12. The first beta-lactamase-producing E. faecium was reported in New Mexico 14. The second report was in a study in Italy in 2011, of which eight isolates were beta-lactamase-producing E. faecium 18. However, beta-lactamase-producing E. faecium is still in scarcity. The relationship between multidrug resistance and biofilm formation is a hypothesis that needs detailed verifications. In our study, VREfa varied adhesin genes and produced robust biofilms. In a study by Donna et al., 72% of isolates carrying esp were in southern California, and 84% were VRE. In another study in Brazil, the esp generated was 72% which was lower than our study 19. In a survey among VRE isolates, the esp gene was observed among 80% of them, nearly consistent with our study 20. In a study, the esp gene was detected among E. faecium isolates from blood, urine, and stools with rates 46%, 38%, and 23%, respectively, significantly lower than our study 12. In a survey by Sharifi et al. among VREfa in northwest Iran, the prevalence of esp gene was 89%, lower than our finding (97%) 21. In Fallah et al., among VREfa from Iran, 75% of them produced biofilms, being significantly higher than ours (56%), and antibiotic resistance in the biofilm formation strains was consistent with the findings of this study. Further verifications did not demonstrate any significant correlation between biofilm formation and esp, asa1 and ebpR genes 23. The rates of esp, asa1 and gelE genes were also 53%, 20% and 51%, respectively, with a lower prevalence of esp and 75% biofilm production, which were more than reported in our study 25. Selected genes involved in the formation of biofilms in other parts of the world were also present in resistant strains with higher resistant rates than biofilm non-producing strains 12,14,16,18-20. Noticeably, the relationship between biofilm formation and antibiotic resistance in Enterococci was confirmed in this study.
CONCLUSION
The antibiotic resistance rate of E. faecium was low. Biofilm formation was significantly different between VREfa and vancomycin-susceptible isolates but not between ampicillin-resistant and ampicillin-susceptible isolates. All the VREfa harbored the ace, esp and ebp genes. The virulence adhesin genes were not significantly associated with biofilm formation. However, there was a significant relation among VREfa that carried all these genes and formed robust biofilms. Further studies are essential for a better understanding of biofilm formation, drug non-susceptibility and adhesin genes.
Funding: This study was not supported by the official site.
Informed Consent Statement: Not applicable.
Data Availability Statement: The results of this study will be found under the journal's rules and related links.
Acknowledgments: To all colleagues in laboratories inside our university.
Conflict of interest: None to declare
REFERENCES
1. Abat, C., D. Raoult, and J.-M.J.M.d.r. Rolain, Low level of resistance in enterococci isolated in four hospitals, Marseille, France. 2016. 22(3): p. 218-222.
2. Ahmed, M.O. and KEJMDR Baptiste, Vancomycin-resistant enterococci: a review of antimicrobial resistance mechanisms and perspectives of human and animal health. 2018. 24(5): p. 590-606.
3. Arias, C.A. and BEJNRM Murray, The rise of the Enterococcus: beyond vancomycin resistance. 2012. 10(4): p. 266-278.
4. Aslangul, E., et al., Acquired gentamicin resistance by permeability impairment in Enterococcus faecalis. 2006. 50(11): p. 3615-3621.
5. Bandyopadhyay, S., et al., prevalence, molecular fingerprinting and drug resistance profile of enterovirulent Escherichia coli isolates from free-ranging yaks of Tawang district, Arunachal Pradesh, India. 2012. 44(5): p. 1063-1072.
6. Mhawesh, A.A., et al., In vitro Experimental Research for Using the Silver Nanoparticles as Plasmid Curing Agent in Some Types of Multi-Antibiotic Resistant Pathogenic Bacteria. 2019. 10(8).
7. Beshiru, A., et al., Multi-antibiotic resistant and putative virulence gene signatures in Enterococcus species isolated from pig farms environment. 2017. 104: p. 90-96.
8. Al-Bdery, A.S.J., G.J. Mohammad, and BJGR Hussen, Vancomycin and linezolid resistance among multidrug-resistant Staphylococcus aureus clinical isolates and interaction with neutrophils. 2020. 21: p. 100804.
9. Ch'ng, J.-H., et al., Biofilm-associated infection by enterococci. 2019. 17(2): p. 82-94.
10. Choi, J.-M. and G.-J.J.C.M. Woo, Transfer of tetracycline resistance genes with aggregation substance in food-borne Enterococcus faecalis. 2015. 70(4): p. 476-484.
11. Wayne, A.J.C.d., Clinical and Laboratory Standards Institute; CLSI. 2011. Performance standards for antimicrobial susceptibility testing. 20th Informational Supplement. 2017.
12. Dale, J.L., et al., Multiple roles for Enterococcus faecalis glycosyltransferases in biofilm-associated antibiotic resistance, cell envelope integrity, and conjugative transfer. 2015. 59(7): p. 4094-4105.
13. Ghasemian, A., et al., expression of adhesin genes and biofilm formation among Klebsiella oxytoca clinical isolates from patients with antibiotic-associated haemorrhagic colitis. 2019. 68(7): p. 978-985.
14. de Jong, A., et al., Antimicrobial susceptibility of enterococci recovered from healthy cattle, pigs and chickens in nine EU countries (EASSA Study) to critically important antibiotics. 2018. 216: p. 168-175.
15. Bhatty, M., et al., E nterococcus faecalis pCF 10‐encoded surface proteins PrgA, PrgB (aggregation substance) and PrgC contribute to plasmid transfer, biofilm formation and virulence. 2015. 95(4): p. 660-677.
16. da Silva Fernandes, M., et al., formation of multi-species biofilms by Enterococcus faecium, Enterococcus faecalis, and Bacillus cereus isolated from ricotta processing and effectiveness of chemical sanitation procedures. 2017. 72: p. 23-28.
17. Wayne, P., Clinical and laboratory standards institute. Performance standards for antimicrobial susceptibility testing. 2011.
18. de Jong, A., et al., Antimicrobial resistance monitoring in commensal enterococci from healthy cattle, pigs and chickens across Europe during 2004–14 (EASSA Study). 2019. 74(4): p. 921-930.
19. Diarra, MS, et al., Distribution of antimicrobial resistance and virulence genes in Enterococcus spp. and characterization of isolates from broiler chickens. 2010. 76(24): p. 8033-8043.
20. Duez, C., et al., The penicillin resistance of Enterococcus faecalis JH2-2r results from an overproduction of the low-affinity penicillin-binding protein PBP4 and does not involve a psr-like geneThe GenBank accession numbers for the sequences reported in this paper are Y17797 for the 8· 4 kb segment of pDML521; AJ290435 for pbp4 of E. faecalis JH2-2; AJ276231 and AJ276232 for the psr-like gene of E. faecalis JH2-2 or JH2-2r, respectively. 2001. 147(9): p. 2561-2569.
21. Duprè, I., et al., Incidence of virulence determinants in clinical Enterococcus faecium and Enterococcus faecalis isolates collected in Sardinia (Italy). 2003. 52(6): p. 491-498.
22. Fair, RJ and Y.J.P.i.m.c. Tor, Antibiotics and bacterial resistance in the 21st century. 2014. 6: p. PMC. S14459.
23. Dworniczek, E., et al., Enterococcus in wound infections: virulence and antimicrobial resistance. 2012. 59(2): p. 263-269.
24. Gök, Ş.M., et al., Investigation of antibiotic resistance and virulence factors of Enterococcus faecium and Enterococcus faecalis strains isolated from clinical samples. 2020. 54(1): p. 26-39.
25. Eaton, T.J., MJJA Gasson, and e. microbiology, Molecular screening of Enterococcus virulence determinants and potential for genetic exchange between food and medical isolates. 2001. 67(4): p. 1628-1635.
Received: 1 December 2021 / Accepted: 9 March 2022 / Published:15 May 2022
Citation: Abeer J. Hassan , Sarmad Ajeel Hazzaa and Dunya Najim Alden Ahmed . Vitamin D a predictor Marker to kidney disease in male with type 2 diabetes mellitus . Revis Bionatura 2022;7(2) 39. http://dx.doi.org/10.21931/RB/2022.07.02.39
